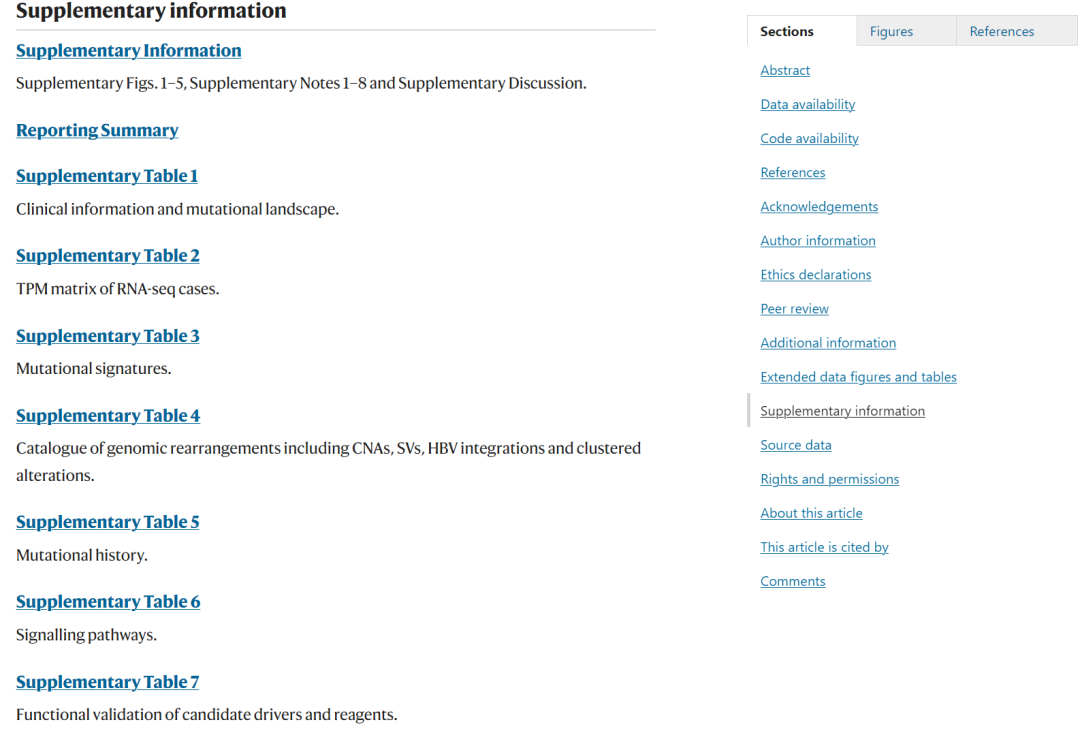
中国人肝癌全基因组项目部分图表重现
项目简介
前面推文介绍过文章 Deep whole-genome analysis of 494 hepatocellular carcinomas,详情见:中国人肝癌全基因组项目。
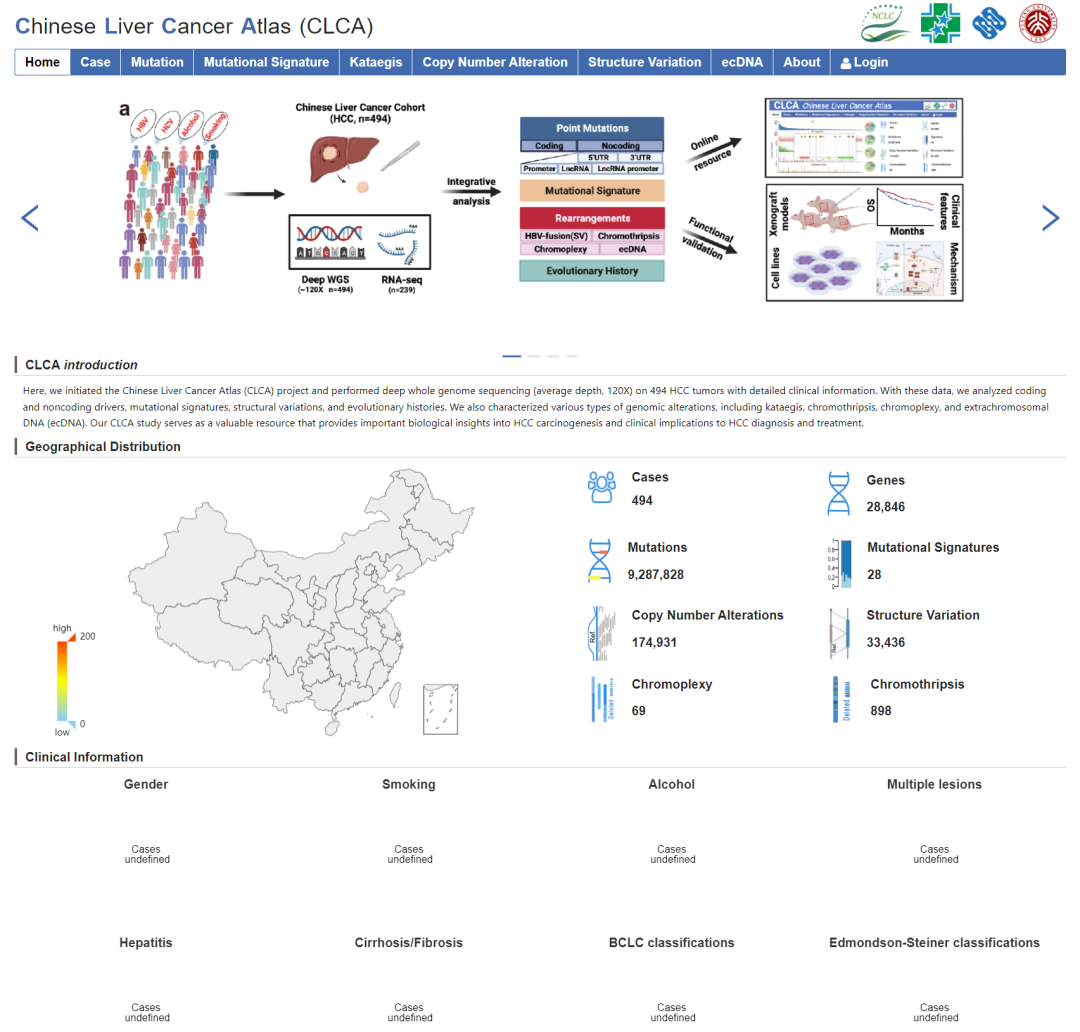
该项目包含494个肝癌病人的 WGS分析结果。作者在文章附件上传了部分数据,同时构建了网页数据库供读者使用。因对文章结果感兴趣,因此从文章附件和网页数据库:http://lifeome.net:8080/clca/#/下载了部分数据进行文章图表重现,数据包括:病人的临床信息、体细胞突变结果,突变特征、拷贝数变异、结构变异、ecDNA等。因为方法上的差异,所以重现结果无法做到和原文一致,如有差异,请以原文分析结果为准。
数据处理
数据下载
这次重现数据来自于文章附件和网页数据库,无需注册登录即可直接下载,很方便:
临床信息
从数据库下载到的临床信息,有 494 个患者,相关的信息有:Province、 Gender、 BCLC、 Age、 Hepatitis、 Cirrhosis/Fibrosis、 Edmondson、 Smoking、 Alcohol、 Multiple、 lesions、 Recurrence、 Death,前 20 位患者的临床信息如下表所示:
# 情况环境并载入R包
rm(list = ls())
library(maftools)
library(stringr)
library(ggpubr)
library(tidyr)
library(data.table)
library(pheatmap)
library(ggrepel)
library(ggsci)
library(ggplot2)
library(VennDiagram)
library(ggVennDiagram)
clinical = rio::import("Cases_20240315.xlsx")
head(clinical,n=20)
CaseID | Province | Gender | BCLC | Age | Hepatitis | Cirrhosis/Fibrosis | Edmondson | Smoking | Alcohol | Multiple lesions | Recurrence | Death |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
CLCA_0001 | Fujian | Male | A | 63 | HBV | Cirrhosis | Level III | No | No | No | No | No |
CLCA_0002 | Henan | Female | A | 76 | HBV | Cirrhosis | Level III | No | No | No | No | No |
CLCA_0003 | Jiangsu | Male | C | 61 | HBV | Cirrhosis | Level III | Yes | Yes | No | Yes | Not Available |
CLCA_0004 | Zhejiang | Male | A | 66 | HBV | Cirrhosis | Level III | Yes | Yes | No | Not Available | Not Available |
CLCA_0005 | Jiangsu | Male | B | 74 | HBV | Fibrosis | Level III | No | No | No | No | No |
CLCA_0006 | Jiangxi | Male | B | 65 | HBV | Fibrosis | Level III | No | No | No | No | No |
CLCA_0007 | Zhejiang | Male | B | 68 | HBV | Cirrhosis | Level II | Yes | No | No | Not Available | Not Available |
CLCA_0008 | Jiangsu | Male | C | 66 | HBV | Cirrhosis | Level III | Yes | Yes | Yes | Yes | Yes |
CLCA_0009 | Jiangsu | Male | B | 69 | HBV | Cirrhosis | Level III | Yes | Yes | Yes | No | No |
CLCA_0010 | Zhejiang | Male | 0 | 65 | HBV | Fibrosis | Level III | No | No | No | Yes | No |
CLCA_0011 | Liaoning | Male | B | 64 | HBV | Fibrosis | Level III | Yes | No | No | Not Available | Not Available |
CLCA_0012 | Anhui | Male | B | 74 | HBV | Fibrosis | Level III | No | No | No | Yes | No |
CLCA_0013 | Fujian | Male | C | 57 | HBV | Fibrosis | Level III | Yes | Yes | Yes | Not Available | Not Available |
CLCA_0014 | Jiangsu | Male | C | 70 | HBV | Fibrosis | Level III | Yes | No | No | Yes | Yes |
CLCA_0015 | Anhui | Male | C | 49 | HBV | Cirrhosis | Level III | Yes | No | No | Not Available | Not Available |
CLCA_0016 | Jiangsu | Male | A | 47 | HBV | Fibrosis | Level III | No | No | No | No | No |
CLCA_0017 | Fujian | Male | C | 61 | HBV | Fibrosis | Level III | No | No | Yes | Not Available | Not Available |
CLCA_0018 | Jiangsu | Male | B | 60 | HBV | Cirrhosis | Level III | Yes | Yes | Yes | Not Available | Not Available |
CLCA_0019 | Jiangxi | Male | B | 79 | HBV | Cirrhosis | Level II | No | No | No | No | No |
CLCA_0020 | Zhejiang | Male | 0 | 56 | HBV | Cirrhosis | Level III | No | No | No | Yes | No |
突变信息
虽然文章中提到鉴定出来的突变有 9287828个,但下载得到的突变信息 Excel表格(可以简单处理为maf格式),显示的也只有 283223个突变位点,这个比例约为3%。因为上传的是注释后的结果,作者WGS 得到的 9287828个突变位点,有很多是落在非编码区或者未知的区域的,只有 283223个约3%的突变位点可以被注释到。
somatic = rio::import("Mutations_20240314.xlsx")
head(somatic,n=20)
CaseID | Gene | Chr | Start | End | Strand | Classification | Type | Ref | Allele | RefReads | AlleleReads | c.HGVS | p.HGVS | transcript |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
CLCA_0001 | RNF223 | chr1 | 1006750 | 1006750 | 3'UTR | SNP | A | G | 143 | 18 | . | . | . | |
CLCA_0001 | PRKCZ | chr1 | 2062535 | 2062535 | promoter | SNP | T | C | 129 | 45 | . | . | . | |
CLCA_0001 | PRKCZ | chr1 | 2103770 | 2103770 | nonsynonymous SNV | SNP | A | T | 123 | 56 | c.1228A>T | p.T410S | NM_002744 | |
CLCA_0001 | LINC00982 | chr1 | 2978977 | 2978977 | lncRNA | SNP | A | T | 194 | 19 | . | . | . | |
CLCA_0001 | PRDM16 | chr1 | 3352980 | 3352980 | 3'UTR | SNP | G | A | 161 | 79 | . | . | . | |
CLCA_0001 | LINC01134 | chr1 | 3831499 | 3831499 | lncRNA | SNP | A | T | 129 | 56 | . | . | . | |
CLCA_0001 | AJAP1 | chr1 | 4849691 | 4849691 | 3'UTR | SNP | A | T | 178 | 60 | . | . | . | |
CLCA_0001 | CHD5 | chr1 | 6163698 | 6163698 | 3'UTR | SNP | A | T | 173 | 17 | . | . | . | |
CLCA_0001 | ICMT | chr1 | 6293565 | 6293565 | nonsynonymous SNV | SNP | T | A | 136 | 49 | c.423A>T | p.L141F | NM_012405 | |
CLCA_0001 | HES2 | chr1 | 6472873 | 6472873 | 3'UTR | SNP | T | A | 167 | 8 | . | . | . | |
CLCA_0001 | HES2 | chr1 | 6476021 | 6476021 | 3'UTR | SNP | A | T | 149 | 52 | . | . | . | |
CLCA_0001 | ESPN | chr1 | 6520683 | 6520683 | 3'UTR | SNP | C | T | 103 | 39 | . | . | . | |
CLCA_0001 | LOC102725193 | chr1 | 7449404 | 7449404 | lncRNA | SNP | C | T | 195 | 82 | . | . | . | |
CLCA_0001 | RERE | chr1 | 8418104 | 8418104 | promoter | SNP | C | A | 202 | 10 | . | . | . | |
CLCA_0001 | G000447 | chr1 | 9217104 | 9217104 | lncRNA | SNP | T | A | 199 | 15 | . | . | . | |
CLCA_0001 | H6PD | chr1 | 9323798 | 9323798 | nonsynonymous SNV | SNP | A | T | 136 | 59 | c.1246A>T | p.R416W | NM_004285 | |
CLCA_0001 | H6PD | chr1 | 9324512 | 9324512 | nonsynonymous SNV | SNP | A | T | 126 | 57 | c.1960A>T | p.M654L | NM_004285 | |
CLCA_0001 | NMNAT1 | chr1 | 10042751 | 10042751 | stopgain | SNP | A | T | 148 | 57 | c.832A>T | p.K278* | NM_022787 | |
CLCA_0001 | G000514 | chr1 | 10686025 | 10686025 | lncRNA | SNP | A | T | 187 | 20 | . | . | . | |
CLCA_0001 | EXOSC10 | chr1 | 11151098 | 11151098 | nonsynonymous SNV | SNP | G | A | 180 | 11 | c.616C>T | p.P206S | NM_001001998 |
# 但是作者文章中方法部分没有提到注释用到的软件,查看其突变注释分类可以看到并非像 VEP 、ANNOVAR 软件注释的
table(somatic$Classification)
##
## 3'UTR 5'UTR
## 73142 20544
## frameshift deletion frameshift insertion
## 1971 698
## lncRNA lncrna.prom
## 48380 10845
## nonframeshift deletion nonframeshift insertion
## 435 66
## nonframeshift substitution nonsynonymous SNV
## 409 52418
## promoter splicing
## 67674 2349
## startloss stopgain
## 158 4001
## stoploss
## 133
图表重现
突变图谱
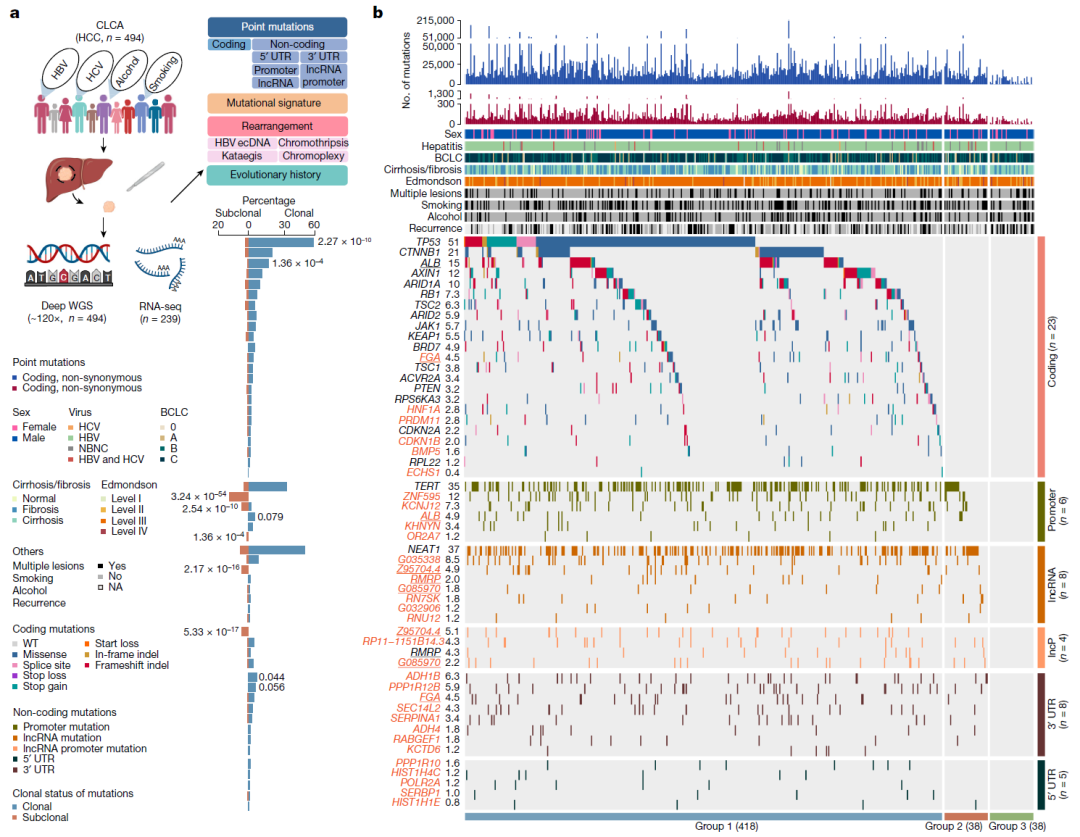
文章中的 fig 1b是体细胞突变图谱,展示的是每个患者特定基因的突变情况,患者有添加上临床信息
# 简单将数据处理一下,以方便后续进行 maftools 处理及可视化
colnames(somatic) = c("Tumor_Sample_Barcode","Hugo_Symbol","Chromosome",
"Start_Position","End_Position","Strand","Variant_Classification",
"Variant_Type","Reference_Allele","Tumor_Seq_Allele2","RefReads","AlleleReads",
"c.HGVS","p.HGVS","transcript")
colnames(clinical)[1] = "Tumor_Sample_Barcode"
# 将临床信息和突变信息读入到 maftools中
maf = read.maf(maf = somatic,vc_nonSyn=unique(somatic$Variant_Classification),clinicalData = clinical)
## -Validating
## --Non MAF specific values in Variant_Classification column:
## promoter
## nonsynonymous SNV
## lncRNA
## stopgain
## splicing
## lncrna.prom
## nonframeshift substitution
## frameshift deletion
## stoploss
## frameshift insertion
## startloss
## nonframeshift deletion
## nonframeshift insertion
## -Summarizing
## --Possible FLAGS among top ten genes:
## TTN
## -Processing clinical data
## -Finished in 7.440s elapsed (49.0s cpu)
# 可以从文章附件中提取到 oncogenes
onco_genes=read.table("onco_genes.txt",header = F)[,1]
# 突变图谱可视化,添加上临床信息
oncoplot(maf,
genes = onco_genes,
keepGeneOrder = T,
annotationFontSize = 1.2,
legendFontSize = 1.0,
removeNonMutated = FALSE,
anno_height = 2,
clinicalFeatures = c("Gender",
"Hepatitis",
"BCLC",
"Cirrhosis/Fibrosis",
"Edmondson",
"Multiple_lesions",
"Smoking",
"Alcohol",
"Recurrence")
)

结果显示只有493名患者,少了一位,但这从下载到的数据就是这样,处理过程并没有改变患者数量,缺失的患者 ID 是 CLCA_0209,从数据库网页下载到的表格中就缺失这个患者的突变信息
sort(unique(somatic$Tumor_Sample_Barcode))
## [1] "CLCA_0001" "CLCA_0002" "CLCA_0003" "CLCA_0004" "CLCA_0005" "CLCA_0006"
## [7] "CLCA_0007" "CLCA_0008" "CLCA_0009" "CLCA_0010" "CLCA_0011" "CLCA_0012"
## [13] "CLCA_0013" "CLCA_0014" "CLCA_0015" "CLCA_0016" "CLCA_0017" "CLCA_0018"
## [19] "CLCA_0019" "CLCA_0020" "CLCA_0021" "CLCA_0022" "CLCA_0023" "CLCA_0024"
## [25] "CLCA_0025" "CLCA_0026" "CLCA_0027" "CLCA_0028" "CLCA_0029" "CLCA_0030"
## [31] "CLCA_0031" "CLCA_0032" "CLCA_0033" "CLCA_0034" "CLCA_0035" "CLCA_0036"
## [37] "CLCA_0037" "CLCA_0038" "CLCA_0039" "CLCA_0040" "CLCA_0041" "CLCA_0042"
## [43] "CLCA_0043" "CLCA_0044" "CLCA_0045" "CLCA_0046" "CLCA_0047" "CLCA_0048"
## [49] "CLCA_0049" "CLCA_0050" "CLCA_0051" "CLCA_0052" "CLCA_0053" "CLCA_0054"
## [55] "CLCA_0055" "CLCA_0056" "CLCA_0057" "CLCA_0058" "CLCA_0059" "CLCA_0060"
## [61] "CLCA_0061" "CLCA_0062" "CLCA_0063" "CLCA_0064" "CLCA_0065" "CLCA_0066"
## [67] "CLCA_0067" "CLCA_0068" "CLCA_0069" "CLCA_0070" "CLCA_0071" "CLCA_0072"
## [73] "CLCA_0073" "CLCA_0074" "CLCA_0075" "CLCA_0076" "CLCA_0077" "CLCA_0078"
## [79] "CLCA_0079" "CLCA_0080" "CLCA_0081" "CLCA_0082" "CLCA_0083" "CLCA_0084"
## [85] "CLCA_0085" "CLCA_0086" "CLCA_0087" "CLCA_0088" "CLCA_0089" "CLCA_0090"
## [91] "CLCA_0091" "CLCA_0092" "CLCA_0093" "CLCA_0094" "CLCA_0095" "CLCA_0096"
## [97] "CLCA_0097" "CLCA_0098" "CLCA_0099" "CLCA_0100" "CLCA_0101" "CLCA_0102"
## [103] "CLCA_0103" "CLCA_0104" "CLCA_0105" "CLCA_0106" "CLCA_0107" "CLCA_0108"
## [109] "CLCA_0109" "CLCA_0110" "CLCA_0111" "CLCA_0112" "CLCA_0113" "CLCA_0114"
## [115] "CLCA_0115" "CLCA_0116" "CLCA_0117" "CLCA_0118" "CLCA_0119" "CLCA_0120"
## [121] "CLCA_0121" "CLCA_0122" "CLCA_0123" "CLCA_0124" "CLCA_0125" "CLCA_0126"
## [127] "CLCA_0127" "CLCA_0128" "CLCA_0129" "CLCA_0130" "CLCA_0131" "CLCA_0132"
## [133] "CLCA_0133" "CLCA_0134" "CLCA_0135" "CLCA_0136" "CLCA_0137" "CLCA_0138"
## [139] "CLCA_0139" "CLCA_0140" "CLCA_0141" "CLCA_0142" "CLCA_0143" "CLCA_0144"
## [145] "CLCA_0145" "CLCA_0146" "CLCA_0147" "CLCA_0148" "CLCA_0149" "CLCA_0150"
## [151] "CLCA_0151" "CLCA_0152" "CLCA_0153" "CLCA_0154" "CLCA_0155" "CLCA_0156"
## [157] "CLCA_0157" "CLCA_0158" "CLCA_0159" "CLCA_0160" "CLCA_0161" "CLCA_0162"
## [163] "CLCA_0163" "CLCA_0164" "CLCA_0165" "CLCA_0166" "CLCA_0167" "CLCA_0168"
## [169] "CLCA_0169" "CLCA_0170" "CLCA_0171" "CLCA_0172" "CLCA_0173" "CLCA_0174"
## [175] "CLCA_0175" "CLCA_0176" "CLCA_0177" "CLCA_0178" "CLCA_0179" "CLCA_0180"
## [181] "CLCA_0181" "CLCA_0182" "CLCA_0183" "CLCA_0184" "CLCA_0185" "CLCA_0186"
## [187] "CLCA_0187" "CLCA_0188" "CLCA_0189" "CLCA_0190" "CLCA_0191" "CLCA_0192"
## [193] "CLCA_0193" "CLCA_0194" "CLCA_0195" "CLCA_0196" "CLCA_0197" "CLCA_0198"
## [199] "CLCA_0199" "CLCA_0200" "CLCA_0201" "CLCA_0202" "CLCA_0203" "CLCA_0204"
## [205] "CLCA_0205" "CLCA_0206" "CLCA_0207" "CLCA_0208" "CLCA_0210" "CLCA_0211"
## [211] "CLCA_0212" "CLCA_0213" "CLCA_0214" "CLCA_0215" "CLCA_0216" "CLCA_0217"
## [217] "CLCA_0218" "CLCA_0219" "CLCA_0220" "CLCA_0221" "CLCA_0222" "CLCA_0223"
## [223] "CLCA_0224" "CLCA_0225" "CLCA_0226" "CLCA_0227" "CLCA_0228" "CLCA_0229"
## [229] "CLCA_0230" "CLCA_0231" "CLCA_0232" "CLCA_0233" "CLCA_0234" "CLCA_0235"
## [235] "CLCA_0236" "CLCA_0237" "CLCA_0238" "CLCA_0239" "CLCA_0240" "CLCA_0241"
## [241] "CLCA_0242" "CLCA_0243" "CLCA_0244" "CLCA_0245" "CLCA_0246" "CLCA_0247"
## [247] "CLCA_0248" "CLCA_0249" "CLCA_0250" "CLCA_0251" "CLCA_0252" "CLCA_0253"
## [253] "CLCA_0254" "CLCA_0255" "CLCA_0256" "CLCA_0257" "CLCA_0258" "CLCA_0259"
## [259] "CLCA_0260" "CLCA_0261" "CLCA_0262" "CLCA_0263" "CLCA_0264" "CLCA_0265"
## [265] "CLCA_0266" "CLCA_0267" "CLCA_0268" "CLCA_0269" "CLCA_0270" "CLCA_0271"
## [271] "CLCA_0272" "CLCA_0273" "CLCA_0274" "CLCA_0275" "CLCA_0276" "CLCA_0277"
## [277] "CLCA_0278" "CLCA_0279" "CLCA_0280" "CLCA_0281" "CLCA_0282" "CLCA_0283"
## [283] "CLCA_0284" "CLCA_0285" "CLCA_0286" "CLCA_0287" "CLCA_0288" "CLCA_0289"
## [289] "CLCA_0290" "CLCA_0291" "CLCA_0292" "CLCA_0293" "CLCA_0294" "CLCA_0295"
## [295] "CLCA_0296" "CLCA_0297" "CLCA_0298" "CLCA_0299" "CLCA_0300" "CLCA_0301"
## [301] "CLCA_0302" "CLCA_0303" "CLCA_0304" "CLCA_0305" "CLCA_0306" "CLCA_0307"
## [307] "CLCA_0308" "CLCA_0309" "CLCA_0310" "CLCA_0311" "CLCA_0312" "CLCA_0313"
## [313] "CLCA_0314" "CLCA_0315" "CLCA_0316" "CLCA_0317" "CLCA_0318" "CLCA_0319"
## [319] "CLCA_0320" "CLCA_0321" "CLCA_0322" "CLCA_0323" "CLCA_0324" "CLCA_0325"
## [325] "CLCA_0326" "CLCA_0327" "CLCA_0328" "CLCA_0329" "CLCA_0330" "CLCA_0331"
## [331] "CLCA_0332" "CLCA_0333" "CLCA_0334" "CLCA_0335" "CLCA_0336" "CLCA_0337"
## [337] "CLCA_0338" "CLCA_0339" "CLCA_0340" "CLCA_0341" "CLCA_0342" "CLCA_0343"
## [343] "CLCA_0344" "CLCA_0345" "CLCA_0346" "CLCA_0347" "CLCA_0348" "CLCA_0349"
## [349] "CLCA_0350" "CLCA_0351" "CLCA_0352" "CLCA_0353" "CLCA_0354" "CLCA_0355"
## [355] "CLCA_0356" "CLCA_0357" "CLCA_0358" "CLCA_0359" "CLCA_0360" "CLCA_0361"
## [361] "CLCA_0362" "CLCA_0363" "CLCA_0364" "CLCA_0365" "CLCA_0366" "CLCA_0367"
## [367] "CLCA_0368" "CLCA_0369" "CLCA_0370" "CLCA_0371" "CLCA_0372" "CLCA_0373"
## [373] "CLCA_0374" "CLCA_0375" "CLCA_0376" "CLCA_0377" "CLCA_0378" "CLCA_0379"
## [379] "CLCA_0380" "CLCA_0381" "CLCA_0382" "CLCA_0383" "CLCA_0384" "CLCA_0385"
## [385] "CLCA_0386" "CLCA_0387" "CLCA_0388" "CLCA_0389" "CLCA_0390" "CLCA_0391"
## [391] "CLCA_0392" "CLCA_0393" "CLCA_0394" "CLCA_0395" "CLCA_0396" "CLCA_0397"
## [397] "CLCA_0398" "CLCA_0399" "CLCA_0400" "CLCA_0401" "CLCA_0402" "CLCA_0403"
## [403] "CLCA_0404" "CLCA_0405" "CLCA_0406" "CLCA_0407" "CLCA_0408" "CLCA_0409"
## [409] "CLCA_0410" "CLCA_0411" "CLCA_0412" "CLCA_0413" "CLCA_0414" "CLCA_0415"
## [415] "CLCA_0416" "CLCA_0417" "CLCA_0418" "CLCA_0419" "CLCA_0420" "CLCA_0421"
## [421] "CLCA_0422" "CLCA_0423" "CLCA_0424" "CLCA_0425" "CLCA_0426" "CLCA_0427"
## [427] "CLCA_0428" "CLCA_0429" "CLCA_0430" "CLCA_0431" "CLCA_0432" "CLCA_0433"
## [433] "CLCA_0434" "CLCA_0435" "CLCA_0436" "CLCA_0437" "CLCA_0438" "CLCA_0439"
## [439] "CLCA_0440" "CLCA_0441" "CLCA_0442" "CLCA_0443" "CLCA_0444" "CLCA_0445"
## [445] "CLCA_0446" "CLCA_0447" "CLCA_0448" "CLCA_0449" "CLCA_0450" "CLCA_0451"
## [451] "CLCA_0452" "CLCA_0453" "CLCA_0454" "CLCA_0455" "CLCA_0456" "CLCA_0457"
## [457] "CLCA_0458" "CLCA_0459" "CLCA_0460" "CLCA_0461" "CLCA_0462" "CLCA_0463"
## [463] "CLCA_0464" "CLCA_0465" "CLCA_0466" "CLCA_0467" "CLCA_0468" "CLCA_0469"
## [469] "CLCA_0470" "CLCA_0471" "CLCA_0472" "CLCA_0473" "CLCA_0474" "CLCA_0475"
## [475] "CLCA_0476" "CLCA_0477" "CLCA_0478" "CLCA_0479" "CLCA_0480" "CLCA_0481"
## [481] "CLCA_0482" "CLCA_0483" "CLCA_0484" "CLCA_0485" "CLCA_0486" "CLCA_0487"
## [487] "CLCA_0488" "CLCA_0489" "CLCA_0490" "CLCA_0491" "CLCA_0492" "CLCA_0493"
## [493] "CLCA_0494"
文章中的突变图谱还对患者进行了分组,Group1 是在oncogene 发生coding突变的患者,Group2则为仅发生 synonymous 突变的患者,Group3为在oncogene上未发生突变的患者(在其他基因有发生突变)。结果显示Group1为418人,Group2为39人,Group3为36人,另外前面提到过突变信息缺少一名患者CLCA_0209。
onco_genes_group1 = onco_genes[1:23]
onco_genes_group2 = onco_genes[24:54]
coding_mutations = c("nonsynonymous SNV",
"stopgain",
"splicing",
"nonframeshift substitution",
"frameshift deletion",
"stoploss",
"frameshift insertion",
"startloss",
"nonframeshift deletion",
"nonframeshift insertion"
)
noncoding_mutations = c("3'UTR","5'UTR","lncRNA","lncrna.prom","promoter")
group1.id = unique(somatic[(somatic$Hugo_Symbol %in% onco_genes_group1) & (somatic$Variant_Classification %in% coding_mutations), 1])
group2.id = setdiff(unique(somatic[(somatic$Hugo_Symbol %in% onco_genes_group2) , 1]),group1.id)
group3.id = setdiff(unique(somatic$Tumor_Sample_Barcode), c(group1.id,group2.id))
group.df = data.frame(Tumor_Sample_Barcode = c(group1.id,
group2.id,
group3.id),
Group = c(rep("Group1",times=length(group1.id)),
rep("Group2",times=length(group2.id)),
rep("Group3",times=length(group3.id)))
)
table(group.df$Group)
##
## Group1 Group2 Group3
## 418 39 36
重新做突变图谱可视化加上 Group 分组信息:
clinical = merge(clinical,group.df,by="Tumor_Sample_Barcode")
maf = read.maf(maf = somatic,
vc_nonSyn=unique(somatic$Variant_Classification),
clinicalData = clinical)
## -Validating
## --Non MAF specific values in Variant_Classification column:
## promoter
## nonsynonymous SNV
## lncRNA
## stopgain
## splicing
## lncrna.prom
## nonframeshift substitution
## frameshift deletion
## stoploss
## frameshift insertion
## startloss
## nonframeshift deletion
## nonframeshift insertion
## -Summarizing
## --Possible FLAGS among top ten genes:
## TTN
## -Processing clinical data
## -Finished in 8.028s elapsed (52.9s cpu)
# 添加上临床信息
oncoplot(maf,
genes = onco_genes,
keepGeneOrder = T,
sortByAnnotation = T,
annotationFontSize = 1.2,
legendFontSize = 1.0,
removeNonMutated = FALSE,
anno_height = 2,
clinicalFeatures = c("Group",
"Gender",
"Hepatitis",
"BCLC",
"Cirrhosis/Fibrosis",
"Edmondson",
"Multiple_lesions",
"Smoking",
"Alcohol",
"Recurrence")
)

突变特征
作者使用的是 mSigHdp 和 SigProfilerExtractor 包进行突变特征分析:
We used mSigHdp (v.1.1.2) and SigProfilerExtractor from SigProfiler bioinformatics tool suite (v.1.1.0)6 to extract SBS, DBS and ID signatures.For SigProfiler signature extraction, 1,000 iterations were performed (nmf_replicates = 1000). We report only signatures supported by both mSigHdp and SigProfiler.
得到的Signature 结果是:
We identified 17 single-base substitution (SBS), 3 doublet-base substitution (DBS) and 8 small insertion-and-deletion (ID) signatures.
除了正文的 fig2 之外,还有 Extended Data fig2
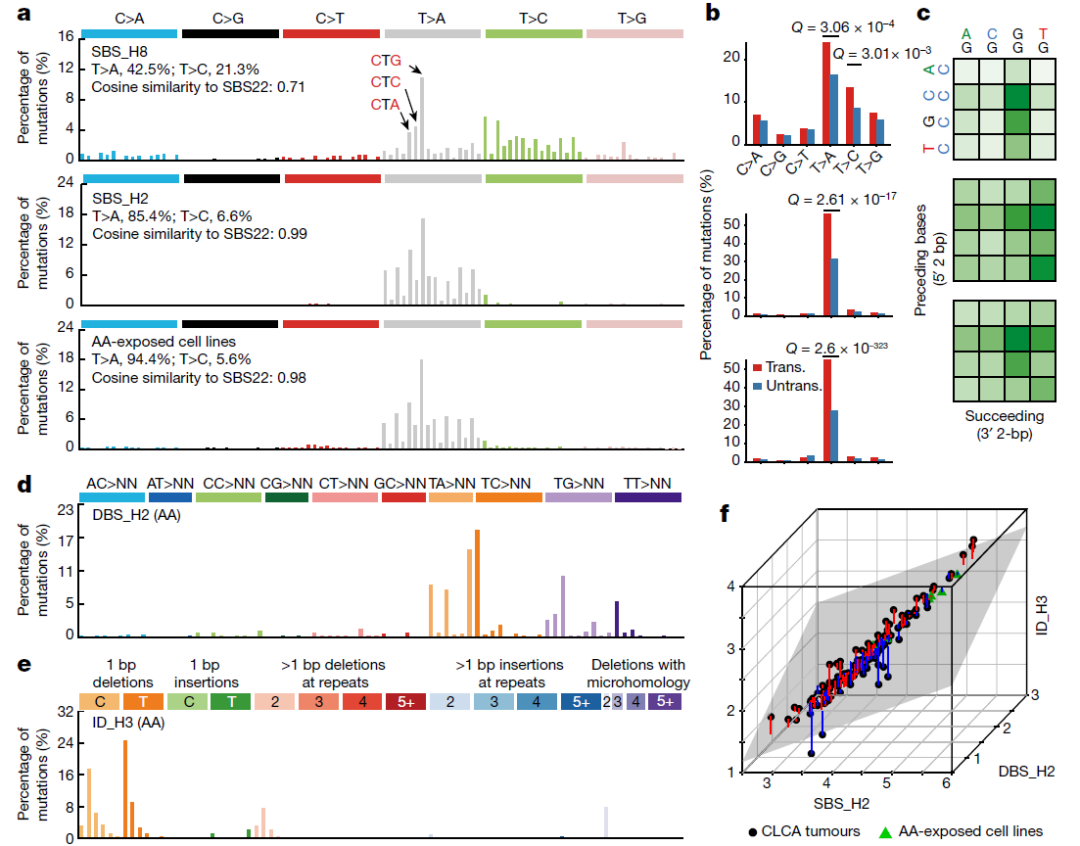

考虑到作者用的方法较为复杂,这里改用maftools 里的signature 分析流程和 sigminer 包的分析流程两种方法:
# 突变特征方法一:maftools ----
library(maftools)
library(NMF)
library(pheatmap)
library(barplot3d)
library(BSgenome.Hsapiens.UCSC.hg19)
# 先构建三连核苷酸矩阵
maf.tnm = trinucleotideMatrix(maf = maf,
#prefix = 'chr',
#add = TRUE,
ref_genome = "BSgenome.Hsapiens.UCSC.hg19")
## -Extracting 5' and 3' adjacent bases
## -Extracting +/- 20bp around mutated bases for background C>T estimation
## -Estimating APOBEC enrichment scores
## --Performing one-way Fisher's test for APOBEC enrichment
## ---APOBEC related mutations are enriched in 0.408 % of samples (APOBEC enrichment score > 2 ; 2 of 490 samples)
## -Creating mutation matrix
## --matrix of dimension 493x96
# 运行 NMF非负矩阵分解,并拟合
# 如果突变较少,需要设置 pConstant = 0.1
maf.sign = estimateSignatures(mat = maf.tnm, nTry = 12)
## -Running NMF for 12 ranks
## Compute NMF rank= 2 ... + measures ... OK
## Compute NMF rank= 3 ... + measures ... OK
## Compute NMF rank= 4 ... + measures ... OK
## Compute NMF rank= 5 ... + measures ... OK
## Compute NMF rank= 6 ... + measures ... OK
## Compute NMF rank= 7 ... + measures ... OK
## Compute NMF rank= 8 ... + measures ... OK
## Compute NMF rank= 9 ... + measures ... OK
## Compute NMF rank= 10 ... + measures ... OK
## Compute NMF rank= 11 ... + measures ... OK
## Compute NMF rank= 12 ... + measures ... OK
## -Finished in 00:07:04 elapsed (00:01:26 cpu)
# 确定最佳突变特征数量
plotCophenetic(res = maf.sign)

# 使用非负矩阵分解将矩阵分解为n签名
maf.sig = extractSignatures(mat = maf.tnm, n = 5)
# 与 COSMIC 的突变特征比较,计算余弦相似度
maf.v3.cosm = compareSignatures(nmfRes = maf.sig, sig_db = "SBS")
# 热图展示余弦相似度
pheatmap::pheatmap(mat = maf.v3.cosm$cosine_similarities, cluster_rows = FALSE, main = "cosine similarity against validated signatures")

# 可视化突变特征
maftools::plotSignatures(nmfRes = maf.sig, title_size = 1.2, sig_db = "SBS")
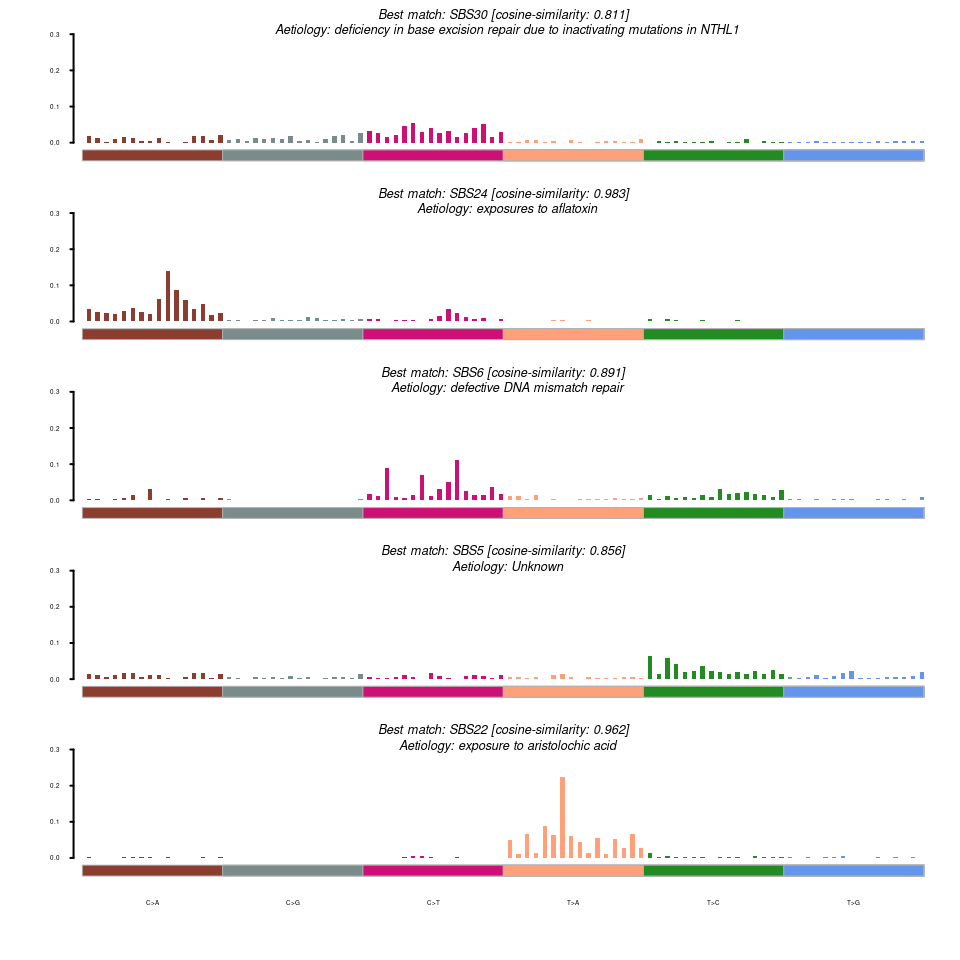
从 maftools 的突变特征分析结果上看,得到的 5 个突变特征分别与 COSMIC 数据库的 SBS30、SBS24、SBS6、SBS5、SBS22 余弦相似度较高。这与原文的结果相差较大,且 maftools 的方法仅分析 SBS 模式的 signature,如果要分析 DBS 或者 INDEL 等 signature,可以使用 sigminer(虽然sigminer 也提供了 SigProfiler的方法,不过用法也相对复杂,这里暂时不考虑。) sigminer 分析的 SBS突变特征有 8个,DBS 有4个,INDEL 有 8个:
# 突变特征方法二:sigminer ----
library(sigminer)
## SBS ----
mt_tally <- sig_tally(
maf,
ref_genome = "BSgenome.Hsapiens.UCSC.hg19",
useSyn = TRUE,
mode = "SBS"
)
mt_sig2 <- sig_unify_extract(mt_tally$nmf_matrix,
range = 10,
nrun = 10)
## 10000 24224.97 25193.74 315481.7 2.800485e-07 8 8
## 10000 24616.58 24754.58 303697.1 3.845836e-06 9 9
## 20000 24616.44 24750.54 303665.8 5.317353e-07 9 9
## 10000 24616.41 24748.37 303630.9 5.924343e-06 9 9
## 20000 24614.28 24739.68 303965.6 2.87272e-05 9 9
## 30000 24612.43 24743.56 304385.7 2.572798e-06 9 9
## 10000 24314.31 25043.34 315276.9 0.0001300736 8 8
## 20000 24294.42 25089.31 316380.3 5.047734e-06 8 8
## 30000 24292.8 25085.5 315789.9 2.433658e-06 8 8
## 40000 24287.52 25093.22 315099.2 1.110607e-05 8 8
## 50000 24284.55 25087.06 314354.7 8.901478e-05 8 8
## 10000 24605.98 24685.06 304196 1.692444e-05 9 9
## 10000 23889.64 25687.55 328564.6 4.943007e-07 7 7
## 10000 24226.84 25185.32 316387.9 1.451832e-07 8 8
## 10000 24604.67 24688.01 303900.7 2.310264e-06 9 9
sim <- get_sig_similarity(mt_sig2, sig_db = "SBS")
pheatmap::pheatmap(sim$similarity)

show_sig_profile(mt_sig2, mode = "SBS", style = "cosmic", x_label_angle = 90)
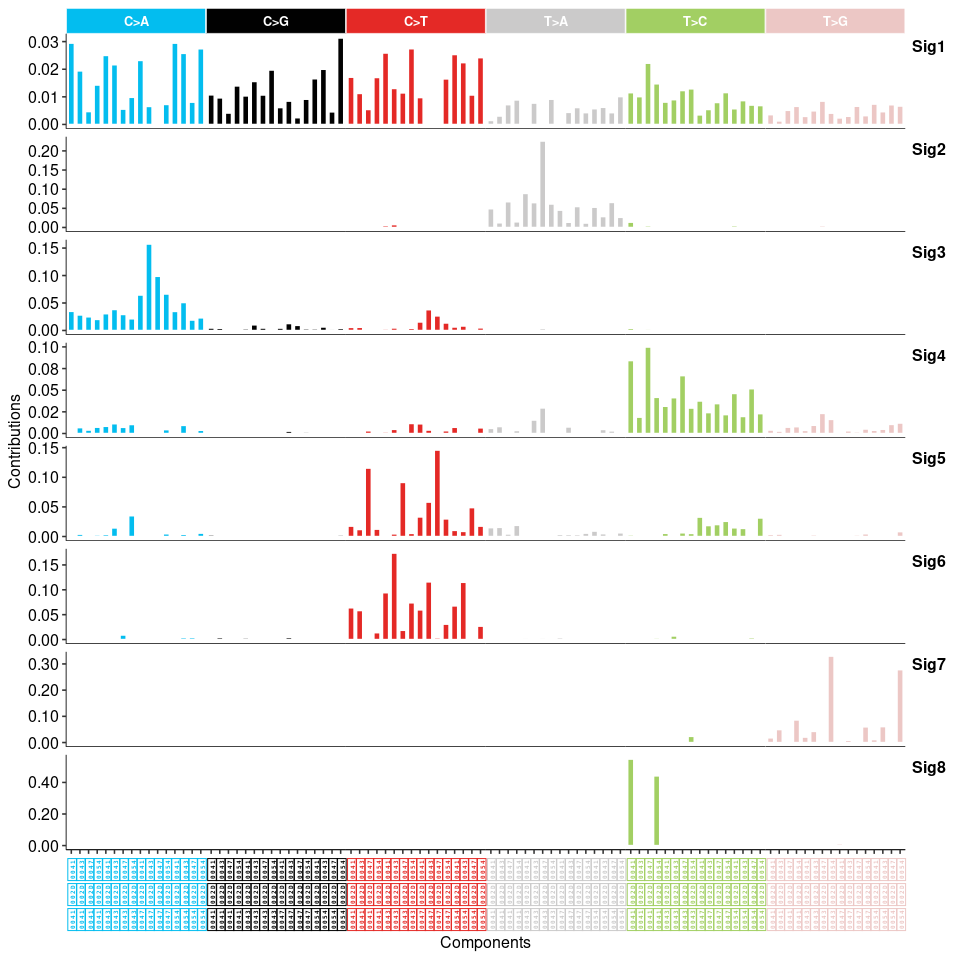
## DBS ----
mt_tally_DBS <- sig_tally(
maf,
ref_genome = "BSgenome.Hsapiens.UCSC.hg19",
useSyn = TRUE,
mode = "DBS"
)
mt_sig2_DBS <- sig_unify_extract(mt_tally_DBS$nmf_matrix,
range = 10,
nrun = 10)
sim_DBS <- get_sig_similarity(mt_sig2_DBS, sig_db = "DBS")
pheatmap::pheatmap(sim_DBS$similarity)
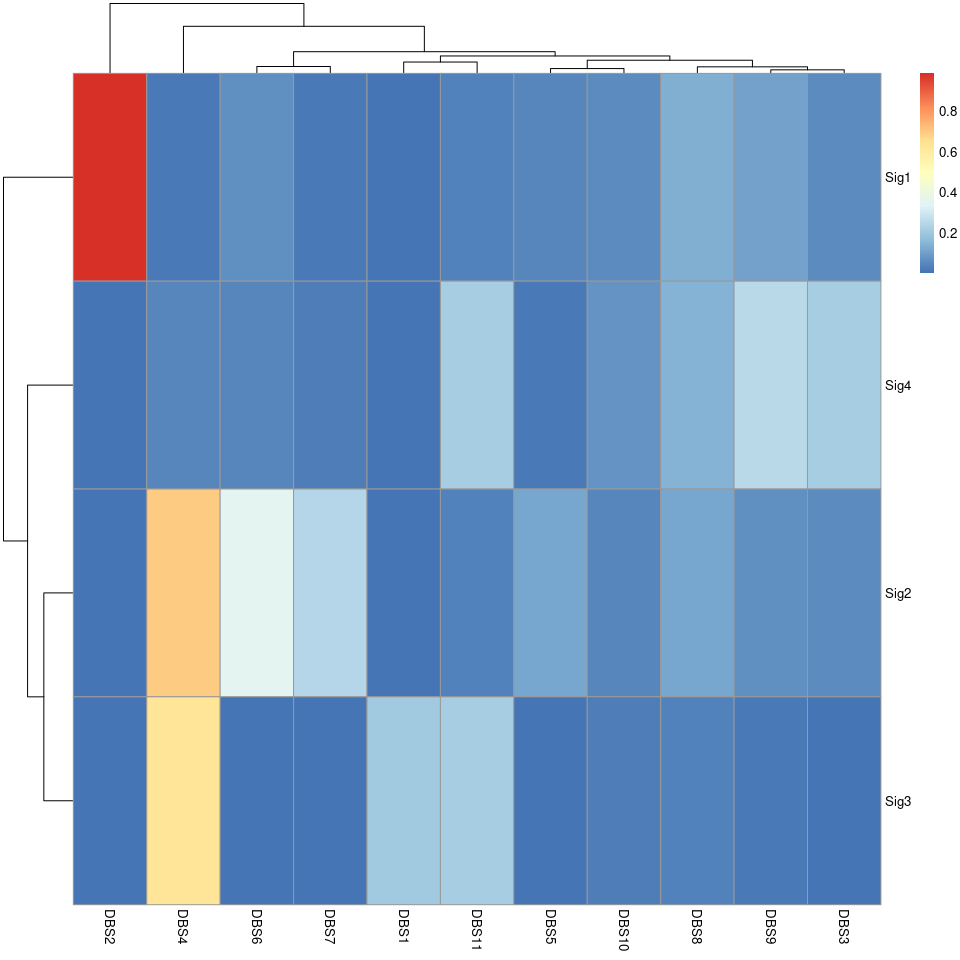
show_sig_profile(mt_sig2_DBS, mode = "DBS", style = "cosmic", x_label_angle = 90)
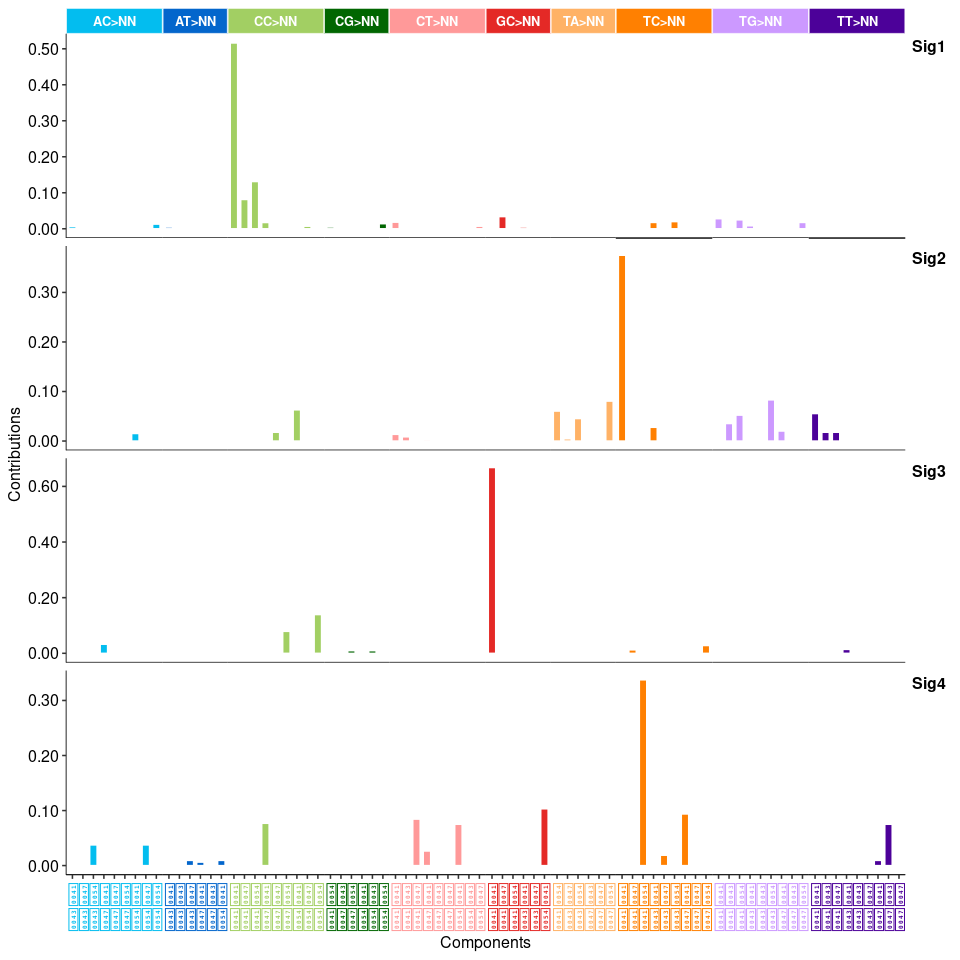
## INDEL ----
mt_tally_ID <- sig_tally(
maf,
ref_genome = "BSgenome.Hsapiens.UCSC.hg19",
useSyn = TRUE,
mode = "ID"
)
mt_sig2_ID <- sig_unify_extract(mt_tally_ID$nmf_matrix,
range = 10,
nrun = 10)
sim_ID <- get_sig_similarity(mt_sig2_ID, sig_db = "ID")
pheatmap::pheatmap(sim_ID$similarity)
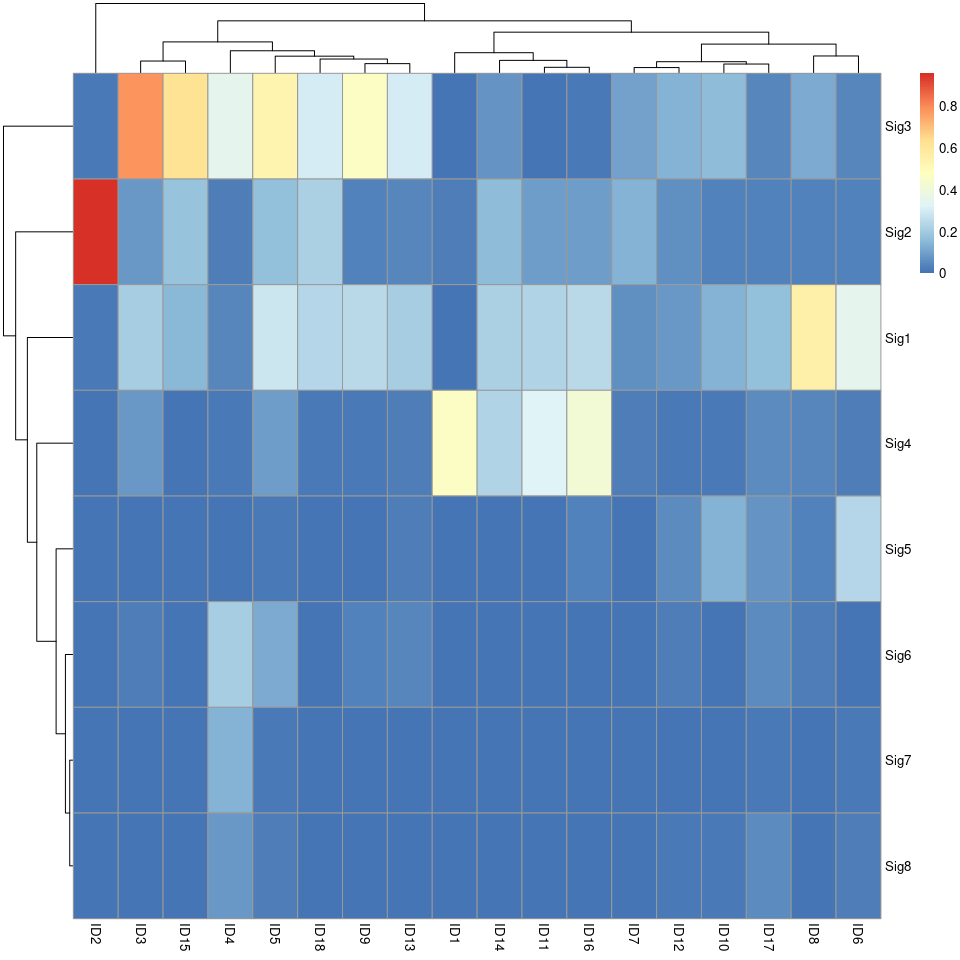
show_sig_profile(mt_sig2_ID, mode = "ID", style = "cosmic", x_label_angle = 90)
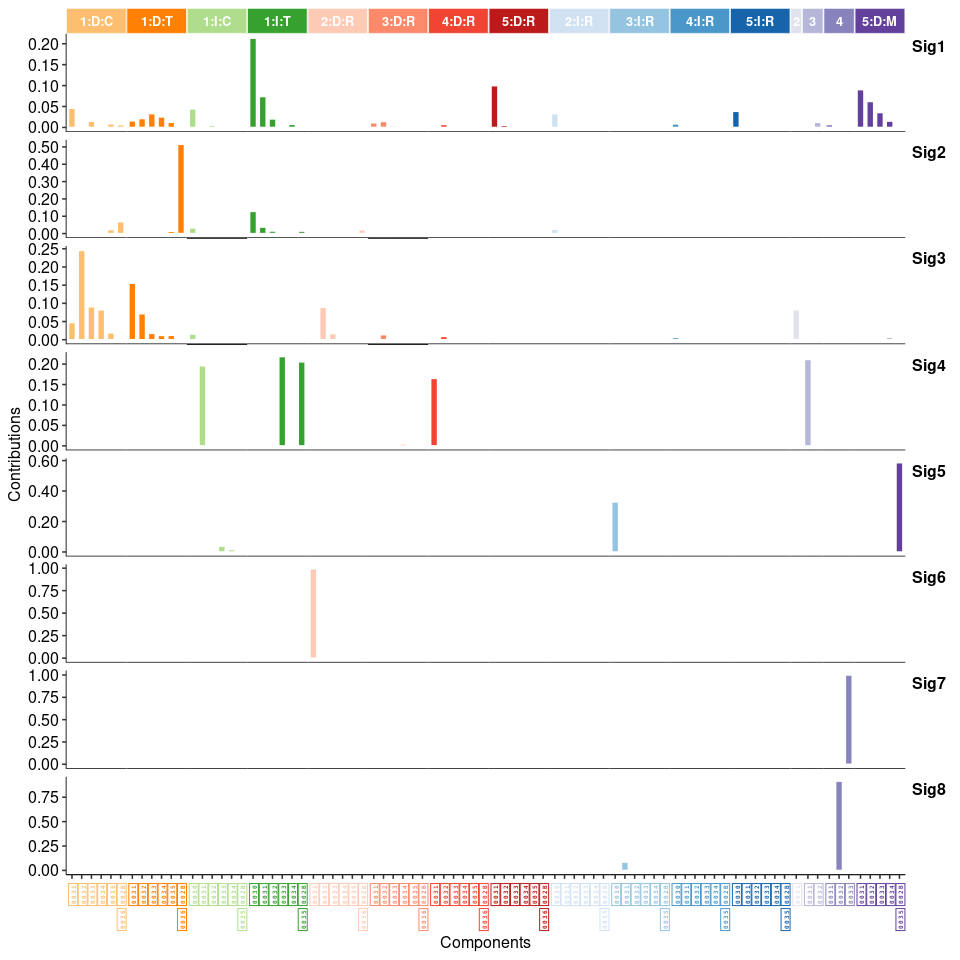
ecDNA 分析
文章的 fig 3 a是ecDNA分析,以饼图形式展示,类型有 BFB、Circular(ecDNA)、Heavily rearranged、Linear 和 No fSCNA 其中 fig3a 原文注释信息是:
The proportion of different amplicons across the CLCA cohort. Circular, breakage–fusion–bridge (BFB), heavily rearranged and linear, and no focal somatic copy-number amplification detected (fSCNA) amplicon categories are shown.
且正文中也提到了:
ecDNA was detected in 27.3% of CLCA tumours
如果这个对应饼图的 Circular(ecDNA) 部分,那就是说在 27.3% 的肿瘤患者中检测到了 ecDNA 事件。
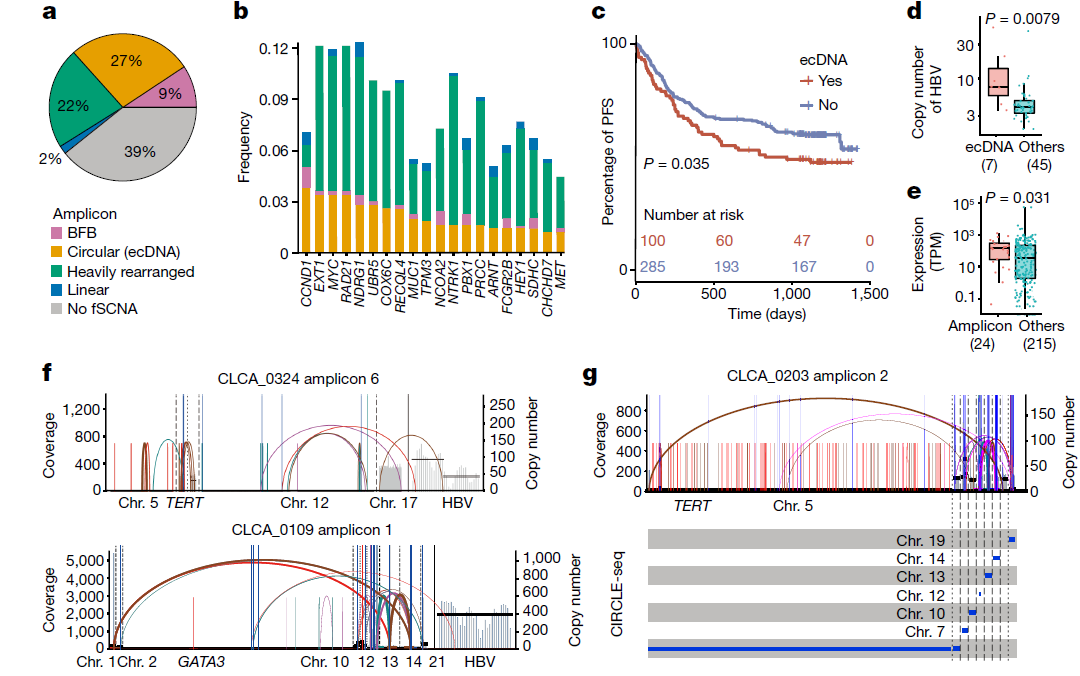
我们可以在文章附件可以找到该图的数据,且数据显示,每一个患者可能发生4种 amp 事件的任意组合 。(注:文章上传的附件Supplementary Table 4:41586_2024_7054_MOESM6_ESM.xlsx 中Table 4g 第一行第三列是 Heavily rearranged rearranged,我的理解应该改为 Heavily rearranged ,以下读入的数据仅手动修改了这一项,其余的没做修改)
amp = readxl::read_xlsx("41586_2024_7054_MOESM6_ESM.xlsx",sheet = 7,skip = 2)
amp = as.data.frame(amp)
# 第一列是患者ID,
# 第二列是amp的类型,
# 第三列是发生某一 amp 类型的 interval counts 数
head(amp,n=100)
sample_name | class | NIntervals | Intervals | OncogenesAmplified | TotalIntervalSize | AmplifiedIntervalSize | AverageAmplifiedCopyCount | Chromosomes | SeqenceEdges | BreakpointEdges | CoverageShifts | MeanshiftSegmentsCopyCount>5 | Foldbacks | CoverageShiftsWithBreakpointEdges |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
CLCA_0001 | Heavily rearranged rearranged | 9 | chr1:179385001-203717000,chr3:129798365-129808957,chr4:9699978-9720571,chr7:5927483-5948075,chr7:6851001-39963000,chr7:62463058-62473650,chr8:18014530-18025122,chr15:40853596-40864189,chr21:33796774-33807367 | ETV1,CDC73,HOXA13,JAZF1,HOXA11,PTPRC,HNRNPA2B1,HOXA9,TPR, | 57538154 | 52083461 | 2.765754199 | 7 | 145 | 59 | 0 | 0 | 0 | 0 |
CLCA_0001 | Heavily rearranged | 4 | chr7:149730451-149741044,chr7:152588001-159138663,chr10:98451408-98662000,chr18:19772106-19792698 | , | 6792443 | 1210371 | 2.635659582 | 3 | 86 | 34 | 1 | 0 | 0 | 1 |
CLCA_0001 | Circular | 10 | chr1:112603401-112813993,chr3:56765044-56775637,chr6:119458107-119568700,chr7:105224001-148714000,chr9:6484724-6695316,chr9:33786290-33806883,chr10:20048538-20059130,chr10:35290516-35311108,chr12:74004366-74014958,chr16:26592312-26612904 | CREB3L2,KIAA1549,POT1,SMO,MET,EZH2,BRAF, | 44115340 | 33806618 | 2.657480486 | 8 | 189 | 77 | 10 | 0 | 2 | 9 |
CLCA_0001 | Heavily rearranged | 10 | chr1:26455415-26466008,chr5:94594394-94604987,chr7:64739205-64749797,chr11:77108686-77119279,chr12:34386996-34413582,chr12:34419001-34560000,chr12:38392853-38603445,chr13:27923699-28134291,chr19:23248743-23259335,chr19:28301468-28342061 | , | 682335 | 177647 | 4.184546951 | 7 | 113 | 25 | 0 | 0 | 0 | 0 |
CLCA_0001 | Heavily rearranged | 5 | chr1:17217822-17238415,chr1:144593226-144603819,chr1:146382400-147865001,chr1:148549086-148559679,chr1:149205805-149246397 | BCL9, | 1564977 | 1074755 | 2.731614502 | 1 | 36 | 13 | 3 | 0 | 0 | 3 |
CLCA_0006 | Circular | 6 | chr4:8438956-8449575,chr6:16220142-16230760,chr7:140356252-140376871,chr10:44079274-44099893,chr11:68389001-69076000,chr19:50449724-50460342 | , | 760098 | 700923 | 6.917366833 | 6 | 35 | 10 | 1 | 0 | 1 | 1 |
CLCA_0008 | Linear | 1 | chr12:1-5343000 | CCND2,KDM5A, | 5343000 | 58827 | 3.867129349 | 1 | 18 | 6 | 3 | 0 | 2 | 3 |
CLCA_0008 | Heavily rearranged | 4 | chr2:179287992-179308657,chr17:46222000-78477201,chr19:28933400-33881601,chrX:48988160-49017173 | CEBPA,CCNE1,CANT1,SRSF2,MSI2,COL1A1,RNF43,DDX5,PRKAR1A,CLTC,BRIP1,HLF,CD79B,H3F3B, | 37253084 | 32087537 | 2.673472586 | 4 | 101 | 37 | 5 | 0 | 0 | 5 |
CLCA_0008 | Heavily rearranged | 5 | chr1:16865110-16885775,chr1:16987734-17018400,chr1:144593611-144614277,chr1:146382400-147845001,chr1:148539013-148559679 | BCL9, | 1555269 | 1143064 | 3.077854662 | 1 | 20 | 5 | 1 | 0 | 0 | 1 |
CLCA_0010 | Heavily rearranged | 10 | chr4:435628-456203,chr5:34438809-34459385,chr5:94594411-94604987,chr6:119458120-119568696,chr7:64873330-64896483,chr8:47908000-146364022,chr10:50452244-50462819,chr12:132926005-132946581,chr18:29064978-29085553,chr19:28319060-28339636 | NCOA2,RECQL4,CHCHD7,EXT1,RAD21,TCEA1,UBR5,NDRG1,MYC,PLAG1,COX6C,HEY1, | 98713790 | 98276040 | 4.541944931 | 9 | 281 | 120 | 8 | 1 | 3 | 8 |
CLCA_0011 | Heavily rearranged | 5 | chr1:17217881-17238420,chr1:144589853-144610392,chr1:146382400-147865001,chr1:148549140-148559679,chr1:149191007-149241546 | BCL9, | 1584762 | 1013046 | 2.739061578 | 1 | 46 | 15 | 2 | 0 | 0 | 2 |
CLCA_0011 | Heavily rearranged | 9 | chr1:112612943-112823482,chr1:204034001-220189000,chr7:35964539-35985077,chr7:62462925-62473464,chr7:116223104-116243643,chr9:6494254-6704793,chr10:20048538-20059076,chr17:39246276-39266814,chr17:39296183-39316721 | MDM4,SLC45A3,ELK4, | 16679316 | 11301097 | 2.574830765 | 5 | 65 | 21 | 6 | 0 | 0 | 6 |
CLCA_0011 | Heavily rearranged | 14 | chr1:4647909-4658448,chr1:156349001-171291000,chr1:174158001-202291000,chr1:226540001-227068000,chr2:117477265-117487804,chr4:437786-455812,chr5:94585136-94605674,chr18:29065444-29085983,chr19:19915204-19965742,chr19:20608823-20639361,chr19:20944620-20955158,chr19:24032644-24043183,chrX:4682464-4693002,chrY:19514391-19524930 | CDC73,PRCC,FCGR2B,NTRK1,PTPRC,TPR,SDHC,ABL2,PBX1, | 43806422 | 12521489 | 2.555923692 | 8 | 147 | 54 | 2 | 0 | 0 | 2 |
CLCA_0011 | Heavily rearranged | 3 | chr1:235674001-242041000,chr5:692512-727516,chr5:766156-796694 | FH, | 6432544 | 60563 | 3.810204351 | 2 | 25 | 12 | 0 | 0 | 0 | 0 |
CLCA_0011 | Heavily rearranged | 2 | chr1:227774001-233912001,chr7:157264846-157275384 | , | 6148540 | 4185741 | 2.542445875 | 2 | 13 | 5 | 0 | 0 | 0 | 0 |
CLCA_0440 | BFB | 1 | chr11:68690001-69655000 | CCND1, | 965000 | 807634 | 11.64473457 | 1 | 15 | 4 | 5 | 1 | 1 | 4 |
CLCA_0440 | Linear | 1 | chr7:77263001-78044000 | , | 781000 | 761473 | 6.945048095 | 1 | 5 | 1 | 1 | 0 | 1 | 1 |
CLCA_0443 | Circular | 7 | chr6:922938-943526,chr6:15337944-15348533,chr8:64227297-64237886,chr8:102644618-102665206,chr11:78306545-78317133,chr12:9841925-9862513,chr13:74163001-115169878 | ERCC5, | 41100414 | 40427161 | 11.58781582 | 5 | 214 | 79 | 52 | 7 | 18 | 48 |
CLCA_0446 | Heavily rearranged | 1 | chr2:195584001-196962000 | , | 1378000 | 1333450 | 3.102743007 | 1 | 19 | 8 | 1 | 0 | 0 | 1 |
CLCA_0446 | Linear | 1 | chr2:81784001-83676000 | , | 1892000 | 1891198 | 3.586781312 | 1 | 35 | 17 | 0 | 0 | 0 | 0 |
CLCA_0446 | BFB | 1 | chr16:48881001-51378000 | CYLD, | 2497000 | 2238057 | 4.281281454 | 1 | 47 | 22 | 3 | 0 | 0 | 2 |
CLCA_0446 | Linear | 1 | chr20:21647001-23960000 | , | 2313000 | 2312969 | 3.797447309 | 1 | 31 | 13 | 1 | 0 | 0 | 1 |
CLCA_0446 | Linear | 1 | chr6:1895001-3964000 | , | 2069000 | 2065337 | 3.474956733 | 1 | 27 | 12 | 0 | 0 | 0 | 0 |
CLCA_0446 | Circular | 2 | chr20:12306001-15355000,chr20:19155447-19165995 | , | 3059549 | 3058791 | 4.059239442 | 1 | 52 | 22 | 2 | 0 | 0 | 1 |
CLCA_0446 | Heavily rearranged | 2 | chr1:170813001-179368000,chr1:222176476-222187024 | ABL2, | 8565549 | 8538860 | 3.531468993 | 1 | 87 | 42 | 0 | 0 | 0 | 0 |
CLCA_0446 | Heavily rearranged | 5 | chr15:20470882-20491430,chr17:50675340-50695889,chr19:28933400-33881601,chr21:28757235-28767784,chrX:48996665-49017214 | CEBPA,CCNE1, | 5020401 | 4958439 | 3.348515983 | 5 | 59 | 24 | 0 | 0 | 0 | 0 |
CLCA_0446 | Heavily rearranged | 6 | chr1:16865098-16885646,chr1:17214199-17244747,chr1:144591196-144603819,chr1:146382400-147866149,chr1:148549130-148569679,chr1:149203321-149233869 | BCL9, | 1598571 | 1412987 | 3.654959179 | 1 | 59 | 21 | 3 | 0 | 0 | 3 |
CLCA_0446 | Heavily rearranged | 9 | chr4:438101-448649,chr5:94594438-94614987,chr7:64739206-64749754,chr7:64873322-64896456,chr12:38490794-38511342,chr17:42089204-42109752,chr18:28342001-29429000,chr19:23258743-23269291,chr19:28293719-28339630 | , | 1249342 | 1215391 | 3.195655085 | 7 | 85 | 25 | 0 | 0 | 0 | 0 |
CLCA_0447 | Circular | 9 | chr1:112703401-112713953,chr2:544014-554566,chr3:8487494-8498046,chr3:137211704-137222256,chr5:37606001-40604000,chr6:32439501-32566760,chr9:1-12739000,chr10:20048538-20059090,chr15:54214001-63803000 | CD274,JAK2,LIFR,TCF12, | 25506025 | 25387775 | 9.946383591 | 8 | 266 | 120 | 22 | 5 | 15 | 16 |
# 每一列的大致信息
str(amp)
## 'data.frame': 2081 obs. of 15 variables:
## $ sample_name : chr "CLCA_0001" "CLCA_0001" "CLCA_0001" "CLCA_0001" ...
## $ class : chr "Heavily rearranged" "Heavily rearranged" "Circular" "Heavily rearranged" ...
## $ NIntervals : num 9 4 10 10 5 6 1 4 5 10 ...
## $ Intervals : chr "chr1:179385001-203717000,chr3:129798365-129808957,chr4:9699978-9720571,chr7:5927483-5948075,chr7:6851001-399630"| __truncated__ "chr7:149730451-149741044,chr7:152588001-159138663,chr10:98451408-98662000,chr18:19772106-19792698" "chr1:112603401-112813993,chr3:56765044-56775637,chr6:119458107-119568700,chr7:105224001-148714000,chr9:6484724-"| __truncated__ "chr1:26455415-26466008,chr5:94594394-94604987,chr7:64739205-64749797,chr11:77108686-77119279,chr12:34386996-344"| __truncated__ ...
## $ OncogenesAmplified : chr "ETV1,CDC73,HOXA13,JAZF1,HOXA11,PTPRC,HNRNPA2B1,HOXA9,TPR," "," "CREB3L2,KIAA1549,POT1,SMO,MET,EZH2,BRAF," "," ...
## $ TotalIntervalSize : num 57538154 6792443 44115340 682335 1564977 ...
## $ AmplifiedIntervalSize : num 52083461 1210371 33806618 177647 1074755 ...
## $ AverageAmplifiedCopyCount : num 2.77 2.64 2.66 4.18 2.73 ...
## $ Chromosomes : num 7 3 8 7 1 6 1 4 1 9 ...
## $ SeqenceEdges : num 145 86 189 113 36 35 18 101 20 281 ...
## $ BreakpointEdges : num 59 34 77 25 13 10 6 37 5 120 ...
## $ CoverageShifts : num 0 1 10 0 3 1 3 5 1 8 ...
## $ MeanshiftSegmentsCopyCount>5 : num 0 0 0 0 0 0 0 0 0 1 ...
## $ Foldbacks : num 0 0 2 0 0 1 2 0 0 3 ...
## $ CoverageShiftsWithBreakpointEdges: num 0 1 9 0 3 1 3 5 1 8 ...
# 总共有 2081 列,amp 信息中,一位患者可以有多行记录,class 类型即为上面提到的类型。
nrow(amp)
## [1] 2081
# 如果直接对表格的第三列 class 进行可视化,会发现结果缺失了 No fSCNA 类型,且比例也不对
library(ggstatsplot)
ggpiestats(
data = amp,
x = class,
palette = "Set1",
#title = "Amplicon",
results.subtitle = F
)
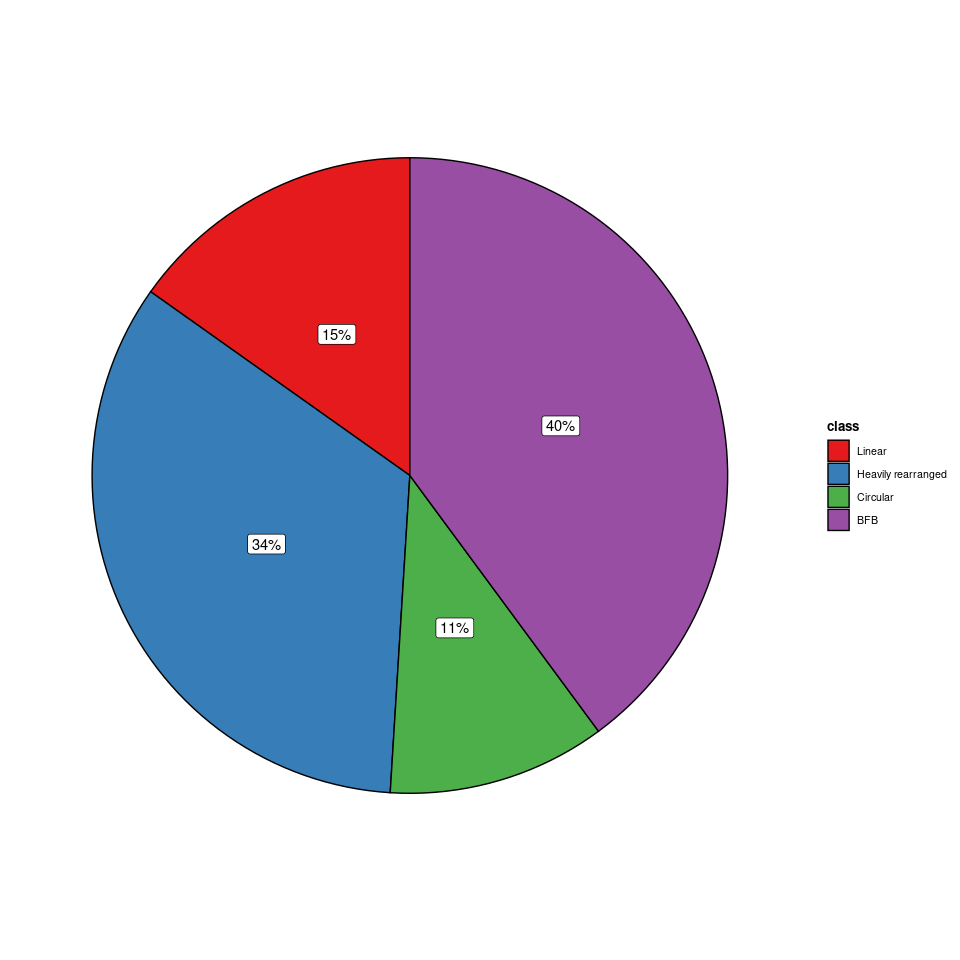
# 这是因为附件的 amp 数据,只包含发生拷贝数变异 Amplicon 的信息,如果患者没有发生,即 No fSCNA 类型,则没有记录在表格中。
table(amp$sample_name)
##
## CLCA_0001 CLCA_0006 CLCA_0008 CLCA_0010 CLCA_0011 CLCA_0013 CLCA_0015 CLCA_0016
## 5 1 3 1 5 7 1 1
## CLCA_0017 CLCA_0018 CLCA_0021 CLCA_0022 CLCA_0023 CLCA_0025 CLCA_0026 CLCA_0029
## 3 6 2 3 2 2 29 3
## CLCA_0031 CLCA_0034 CLCA_0038 CLCA_0039 CLCA_0040 CLCA_0044 CLCA_0045 CLCA_0046
## 6 2 4 4 2 7 3 2
## CLCA_0052 CLCA_0056 CLCA_0059 CLCA_0060 CLCA_0061 CLCA_0062 CLCA_0065 CLCA_0066
## 6 24 5 1 1 5 1 4
## CLCA_0067 CLCA_0068 CLCA_0069 CLCA_0070 CLCA_0071 CLCA_0072 CLCA_0073 CLCA_0074
## 3 3 1 4 4 3 4 7
## CLCA_0078 CLCA_0079 CLCA_0080 CLCA_0089 CLCA_0090 CLCA_0091 CLCA_0092 CLCA_0093
## 2 4 7 5 4 3 3 4
## CLCA_0095 CLCA_0096 CLCA_0097 CLCA_0098 CLCA_0099 CLCA_0100 CLCA_0101 CLCA_0102
## 4 13 8 2 1 4 2 1
## CLCA_0103 CLCA_0104 CLCA_0105 CLCA_0106 CLCA_0107 CLCA_0108 CLCA_0109 CLCA_0110
## 3 2 4 3 2 2 1 1
## CLCA_0111 CLCA_0112 CLCA_0113 CLCA_0114 CLCA_0115 CLCA_0116 CLCA_0118 CLCA_0119
## 1 13 2 9 3 4 149 13
## CLCA_0120 CLCA_0121 CLCA_0122 CLCA_0123 CLCA_0125 CLCA_0126 CLCA_0128 CLCA_0129
## 235 20 31 217 39 1 201 2
## CLCA_0130 CLCA_0132 CLCA_0133 CLCA_0135 CLCA_0137 CLCA_0139 CLCA_0140 CLCA_0141
## 4 5 2 2 1 1 3 5
## CLCA_0143 CLCA_0144 CLCA_0145 CLCA_0146 CLCA_0147 CLCA_0148 CLCA_0150 CLCA_0153
## 7 2 9 2 1 3 8 45
## CLCA_0154 CLCA_0156 CLCA_0157 CLCA_0158 CLCA_0159 CLCA_0160 CLCA_0165 CLCA_0166
## 7 4 2 5 8 1 4 3
## CLCA_0167 CLCA_0168 CLCA_0171 CLCA_0173 CLCA_0174 CLCA_0176 CLCA_0177 CLCA_0178
## 6 5 3 1 1 7 1 3
## CLCA_0182 CLCA_0187 CLCA_0188 CLCA_0189 CLCA_0190 CLCA_0191 CLCA_0192 CLCA_0194
## 5 1 1 1 3 8 3 1
## CLCA_0197 CLCA_0198 CLCA_0201 CLCA_0202 CLCA_0203 CLCA_0204 CLCA_0205 CLCA_0206
## 2 2 4 5 5 2 3 1
## CLCA_0207 CLCA_0208 CLCA_0210 CLCA_0212 CLCA_0215 CLCA_0216 CLCA_0217 CLCA_0218
## 4 1 3 2 7 2 2 5
## CLCA_0219 CLCA_0221 CLCA_0222 CLCA_0223 CLCA_0224 CLCA_0227 CLCA_0229 CLCA_0231
## 4 10 6 8 5 1 2 1
## CLCA_0232 CLCA_0233 CLCA_0235 CLCA_0236 CLCA_0237 CLCA_0239 CLCA_0243 CLCA_0245
## 2 1 2 1 1 6 9 1
## CLCA_0246 CLCA_0248 CLCA_0249 CLCA_0251 CLCA_0254 CLCA_0255 CLCA_0256 CLCA_0257
## 4 1 1 1 1 1 1 1
## CLCA_0258 CLCA_0259 CLCA_0261 CLCA_0263 CLCA_0265 CLCA_0268 CLCA_0270 CLCA_0271
## 2 4 4 3 3 2 2 5
## CLCA_0277 CLCA_0278 CLCA_0281 CLCA_0282 CLCA_0283 CLCA_0284 CLCA_0285 CLCA_0289
## 1 2 1 2 3 2 5 4
## CLCA_0291 CLCA_0293 CLCA_0294 CLCA_0295 CLCA_0296 CLCA_0301 CLCA_0303 CLCA_0305
## 1 1 3 4 1 3 2 1
## CLCA_0309 CLCA_0310 CLCA_0311 CLCA_0314 CLCA_0315 CLCA_0316 CLCA_0317 CLCA_0321
## 1 1 2 2 3 1 3 1
## CLCA_0323 CLCA_0324 CLCA_0325 CLCA_0327 CLCA_0330 CLCA_0331 CLCA_0332 CLCA_0334
## 2 4 11 1 4 4 3 2
## CLCA_0336 CLCA_0337 CLCA_0338 CLCA_0341 CLCA_0342 CLCA_0343 CLCA_0344 CLCA_0345
## 5 5 3 1 2 3 3 4
## CLCA_0346 CLCA_0347 CLCA_0348 CLCA_0349 CLCA_0351 CLCA_0352 CLCA_0354 CLCA_0356
## 2 4 2 3 4 6 1 5
## CLCA_0357 CLCA_0359 CLCA_0365 CLCA_0366 CLCA_0367 CLCA_0369 CLCA_0372 CLCA_0373
## 14 2 3 10 3 11 1 5
## CLCA_0375 CLCA_0376 CLCA_0377 CLCA_0378 CLCA_0379 CLCA_0382 CLCA_0384 CLCA_0385
## 3 1 12 1 2 6 13 4
## CLCA_0387 CLCA_0388 CLCA_0389 CLCA_0390 CLCA_0391 CLCA_0392 CLCA_0393 CLCA_0394
## 1 1 4 20 1 12 4 1
## CLCA_0395 CLCA_0398 CLCA_0399 CLCA_0400 CLCA_0401 CLCA_0402 CLCA_0403 CLCA_0404
## 3 9 1 1 4 4 2 4
## CLCA_0406 CLCA_0407 CLCA_0408 CLCA_0409 CLCA_0410 CLCA_0411 CLCA_0412 CLCA_0413
## 11 5 7 2 2 2 2 1
## CLCA_0414 CLCA_0416 CLCA_0418 CLCA_0419 CLCA_0420 CLCA_0421 CLCA_0424 CLCA_0425
## 1 4 2 10 4 1 4 1
## CLCA_0426 CLCA_0428 CLCA_0429 CLCA_0433 CLCA_0435 CLCA_0439 CLCA_0440 CLCA_0443
## 2 1 14 3 7 2 2 1
## CLCA_0446 CLCA_0447 CLCA_0448 CLCA_0450 CLCA_0451 CLCA_0458 CLCA_0461 CLCA_0462
## 10 18 3 1 7 2 3 1
## CLCA_0465 CLCA_0467 CLCA_0470 CLCA_0472 CLCA_0474 CLCA_0475 CLCA_0477 CLCA_0478
## 5 1 1 3 2 1 16 3
## CLCA_0479 CLCA_0480 CLCA_0481 CLCA_0482 CLCA_0484 CLCA_0485 CLCA_0486 CLCA_0487
## 2 20 7 23 2 9 7 4
## CLCA_0488 CLCA_0492 CLCA_0493 CLCA_0494
## 1 1 3 2
table(amp$class)
##
## BFB Circular Heavily rearranged Linear
## 830 231 704 316
# 总共是494名患者,其中amp 表格记录的患者有 300 名
unique(amp$sample_name) %>% length()
## [1] 300
# 那么没有 amp 记录的患者就是 194 名,比例为 39% 和原图符合
194/494
## [1] 0.3927126
# 先简单粗暴地获取每一种amp类型的患者ID
BFB.id = unique(amp[amp$class == "BFB",1])
Circular.id = unique(amp[amp$class == "Circular",1])
Heavily_rearranged.id = unique(amp[amp$class == "Heavily rearranged",1])
Linear.id = unique(amp[amp$class == "Linear",1])
No_fSCNA.id = setdiff(clinical$Tumor_Sample_Barcode, unique(amp$sample_name))
length(BFB.id);length(Circular.id);length(Heavily_rearranged.id);length(Linear.id);length(No_fSCNA.id)
## [1] 81
## [1] 135
## [1] 233
## [1] 137
## [1] 193
韦恩图进行可视化可以发现,这样获取到的患者ID是有交集的,前面就提到过了,每一个患者可能发生4种 amp 事件的任意组合。所以有交集才是正常的。但这样的话,原文的饼图就无法解释了。
# 韦恩图进行可视化
amp.list = list(BFB.id,Circular.id,Heavily_rearranged.id,Linear.id,No_fSCNA.id)
names(amp.list) = c('BFB','Circular','Heavily_rearranged','Linear','No_fSCNA')
venn.plot1 <- venn.diagram(
x = amp.list,
col = "transparent",
euler.d = TRUE,
fill = c("#E64B35B2", "#4DBBD5B2", "#00A087B2", "#3C5488B2", "#F39B7FB2"),
alpha = rep(0.6,time = 5),
cex = 1.2,
cat.cex = 1.0,
# main = patients[i],
main.cex = 1.0,
print.mode = c("raw", "percent"),
category.names = names(amp.list),
filename = NULL
)
p = as_ggplot(venn.plot1)
print(p)
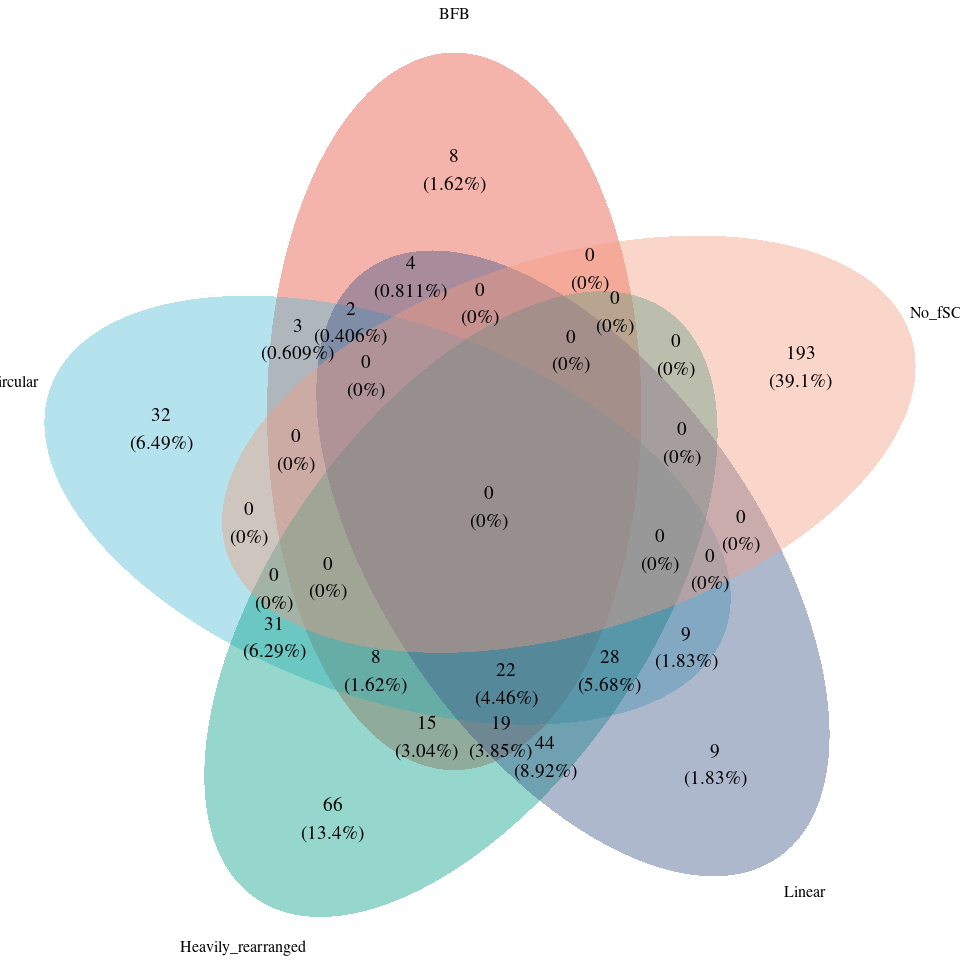
尝试探索一下数据以获取和原文中的比例接近的结果。从数据上看,发生 Circular(ecDNA) 患者是 135名, 135/494=27.3% 符合原文饼图比例。但其他amp事件Heavily rearranged、Linear、BFB 就不符合比例了,不满足。除非取差集,也就是对 amp 事件划分优先级,发生 Circular(ecDNA) 事件的患者不再记录其他事件,即 Circular(ecDNA) > BFB > Heavily rearranged >Linear,这样比例符合了,但无法理解这样做的意义何在?
# Circular(ecDNA)
length(Circular.id)/494
## [1] 0.2732794
# BFB
setdiff(BFB.id,Circular.id) %>% length() /494
## [1] 0.09311741
# Heavily rearranged
setdiff(Heavily_rearranged.id,c(BFB.id,Circular.id)) %>% length() /494
## [1] 0.2226721
# Linear
setdiff(Linear.id,c(BFB.id,Circular.id,Heavily_rearranged.id)) %>% length() /494
## [1] 0.01821862
# 虽然这样结果和作者的结果吻合,但是这样做的意义何在呢?
amp2 = data.frame(sample_name = c(Circular.id,
setdiff(BFB.id,Circular.id),
setdiff(Heavily_rearranged.id,c(BFB.id,Circular.id)),
setdiff(Linear.id,c(BFB.id,Circular.id,Heavily_rearranged.id)),
No_fSCNA.id
),
class = c(rep("Circular",times = length(Circular.id)),
rep("BFB",times = length(setdiff(BFB.id,Circular.id))),
rep("Heavily_rearranged",times = length(setdiff(Heavily_rearranged.id,
c(BFB.id,Circular.id)))),
rep("Linear",times = length(setdiff(Linear.id,
c(BFB.id,
Circular.id,
Heavily_rearranged.id)))),
rep("No_fSCNA",times = length(No_fSCNA.id)))
)
# 饼图
ggpiestats(
data = amp2,
x = class,
palette = "Set1",
#title = "Amplicon",
results.subtitle = F
)
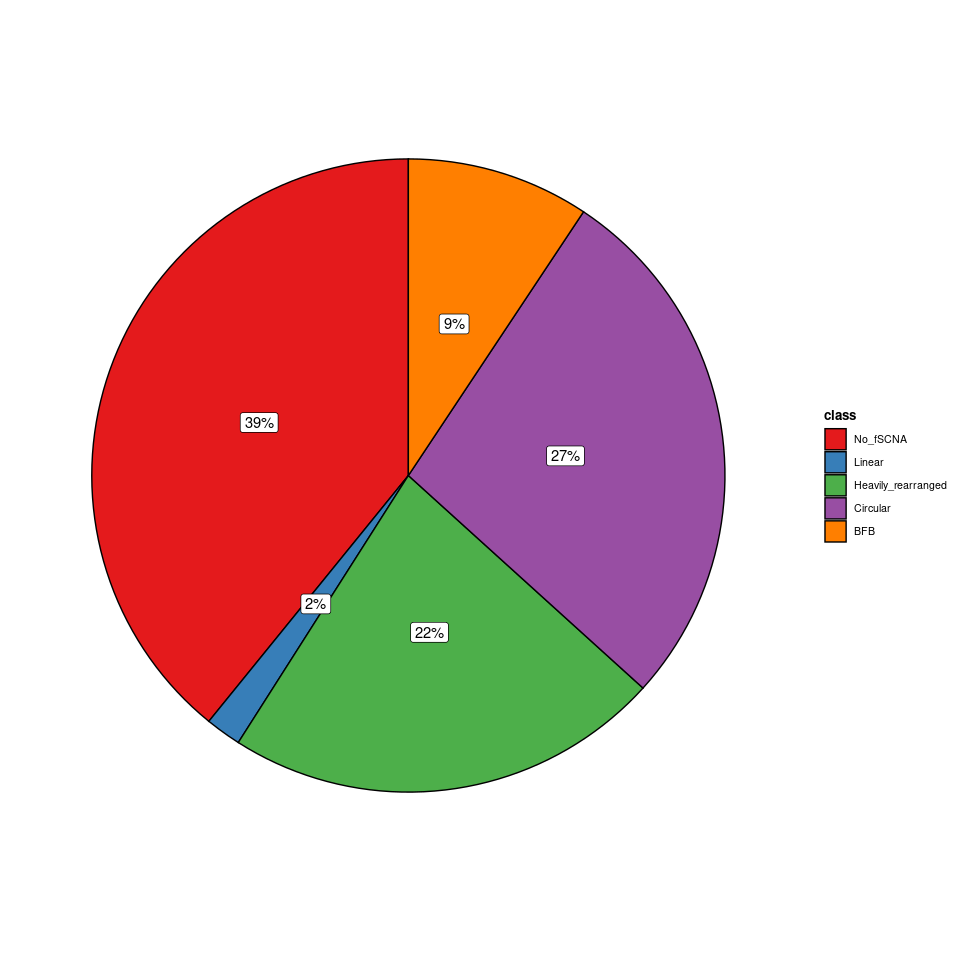
fig3b 是 ecDNA 上的基因列表,进行柱状图可视化。但是根据作者上传的附件重现出来的结果和文章的 fig.3b 并不止一致,如文章原图中的 EXT1 MYC RAD21 NDRG1柱子高度相接近,但上面可视化出来的结果显示MYC 较高,其他的较低。
# 获取 ecDNA
ecDNA_amp = amp[amp$class=="Circular",]
head(ecDNA_amp,n=20)
sample_name | class | NIntervals | Intervals | OncogenesAmplified | TotalIntervalSize | AmplifiedIntervalSize | AverageAmplifiedCopyCount | Chromosomes | SeqenceEdges | BreakpointEdges | CoverageShifts | MeanshiftSegmentsCopyCount>5 | Foldbacks | CoverageShiftsWithBreakpointEdges |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
CLCA_0001 | Circular | 10 | chr1:112603401-112813993,chr3:56765044-56775637,chr6:119458107-119568700,chr7:105224001-148714000,chr9:6484724-6695316,chr9:33786290-33806883,chr10:20048538-20059130,chr10:35290516-35311108,chr12:74004366-74014958,chr16:26592312-26612904 | CREB3L2,KIAA1549,POT1,SMO,MET,EZH2,BRAF, | 44115340 | 33806618 | 2.65748 | 8 | 189 | 77 | 10 | 0 | 2 | 9 |
CLCA_0006 | Circular | 6 | chr4:8438956-8449575,chr6:16220142-16230760,chr7:140356252-140376871,chr10:44079274-44099893,chr11:68389001-69076000,chr19:50449724-50460342 | , | 760098 | 700923 | 6.917367 | 6 | 35 | 10 | 1 | 0 | 1 | 1 |
CLCA_0443 | Circular | 7 | chr6:922938-943526,chr6:15337944-15348533,chr8:64227297-64237886,chr8:102644618-102665206,chr11:78306545-78317133,chr12:9841925-9862513,chr13:74163001-115169878 | ERCC5, | 41100414 | 40427161 | 11.58782 | 5 | 214 | 79 | 52 | 7 | 18 | 48 |
CLCA_0446 | Circular | 2 | chr20:12306001-15355000,chr20:19155447-19165995 | , | 3059549 | 3058791 | 4.059239 | 1 | 52 | 22 | 2 | 0 | 0 | 1 |
CLCA_0447 | Circular | 9 | chr1:112703401-112713953,chr2:544014-554566,chr3:8487494-8498046,chr3:137211704-137222256,chr5:37606001-40604000,chr6:32439501-32566760,chr9:1-12739000,chr10:20048538-20059090,chr15:54214001-63803000 | CD274,JAK2,LIFR,TCF12, | 25506025 | 25387775 | 9.946384 | 8 | 266 | 120 | 22 | 5 | 15 | 16 |
CLCA_0447 | Circular | 1 | chr3:171543001-172009000 | , | 466000 | 446113 | 4.344856 | 1 | 6 | 1 | 2 | 0 | 1 | 2 |
CLCA_0447 | Circular | 1 | chr22:35836001-36188000 | , | 352000 | 351993 | 3.970767 | 1 | 7 | 3 | 0 | 0 | 0 | 0 |
CLCA_0447 | Circular | 1 | chr15:65816001-66021000 | , | 205000 | 204997 | 4.891946 | 1 | 3 | 1 | 0 | 0 | 0 | 0 |
CLCA_0447 | Circular | 1 | chr6:43482001-44354000 | , | 872000 | 852707 | 3.857693 | 1 | 9 | 2 | 1 | 0 | 0 | 1 |
CLCA_0447 | Circular | 1 | chr15:67144001-67445000 | , | 301000 | 300105 | 5.693422 | 1 | 6 | 2 | 0 | 0 | 0 | 0 |
CLCA_0451 | Circular | 7 | chr2:70504051-70524611,chr5:112940001-118170000,chr6:32432533-32579209,chr7:40448001-43603000,chr8:109545001-120080000,chr11:113948001-116387000,chr11:130256001-135006516 | EXT1,RAD21, | 26276754 | 41610 | 3.206914 | 6 | 209 | 92 | 1 | 0 | 0 | 1 |
CLCA_0461 | Circular | 16 | chr1:4649294-4659888,chr1:150319900-223199001,chr1:225205000-233912001,chr4:437018-457611,chr5:40686831-40697425,chr5:94594161-94604755,chr7:64865863-64896457,chr12:38490794-38511387,chr17:42089772-42100365,chr18:29073967-29084561,chr19:20599211-20629805,chr19:20944362-20955213,chr19:23476178-23486772,chr19:28280720-28349688,chrX:4689289-4699883,chrY:19505241-19515835 | H3F3A,ARNT,PRCC,FCGR2B,MUC1,CDC73,TPM3,NTRK1,SLC45A3,PTPRC,TPR,ELK4,SDHC,ABL2,MDM4,PBX1, | 81853062 | 81341590 | 4.435998 | 10 | 366 | 127 | 10 | 0 | 0 | 9 |
CLCA_0465 | Circular | 4 | chr11:2189298-2209845,chr11:59494001-59723000,chr11:60340001-61377000,chr11:68706001-70495000 | CCND1, | 3075548 | 2898925 | 4.283449 | 1 | 37 | 10 | 9 | 0 | 2 | 7 |
CLCA_0465 | Circular | 8 | chr5:34438837-34459385,chr6:32432537-32579209,chr6:119458160-119658708,chr8:69214001-146364022,chr10:50452244-50462791,chr12:131860272-131880819,chr12:132926033-132936581,chr20:1380675-1391222 | NCOA2,RECQL4,EXT1,RAD21,COX6C,NDRG1,MYC,UBR5,HEY1, | 77569986 | 76966860 | 3.3338 | 6 | 376 | 156 | 9 | 1 | 2 | 8 |
CLCA_0470 | Circular | 1 | chr1:154834001-155367000 | MUC1, | 533000 | 532996 | 5.465167 | 1 | 4 | 2 | 0 | 0 | 0 | 0 |
CLCA_0472 | Circular | 5 | chr16:46518719-46529341,chr16:46552064-46649972,chr16:46715036-46735657,chr16:46767016-46787638,chr16:46825001-49898000 | , | 3222777 | 3200819 | 7.860472 | 1 | 100 | 43 | 8 | 0 | 0 | 7 |
CLCA_0480 | Circular | 3 | chr6:32434297-32464893,chr6:32478990-32571656,chr20:832001-2745000 | , | 2036264 | 32725 | 3.50202 | 2 | 94 | 45 | 0 | 0 | 0 | 0 |
CLCA_0481 | Circular | 10 | chr1:112603401-112814054,chr3:197842684-197853337,chr4:190896293-190916947,chr5:34438730-34459384,chr6:119458046-119668700,chr7:116222989-116233643,chr9:6494140-6604794,chr10:18594001-37659000,chr16:26592312-26612965,chr18:14772319-14782973 | KIF5B,ABI1,MLLT10, | 19690892 | 16577851 | 2.984219 | 10 | 151 | 56 | 6 | 0 | 0 | 6 |
CLCA_0482 | Circular | 1 | chr1:196437001-197013000 | , | 576000 | 471329 | 3.063173 | 1 | 19 | 7 | 4 | 0 | 0 | 4 |
CLCA_0484 | Circular | 4 | chr10:1746001-4002667,chr11:66975001-67656269,chr11:71568859-71579471,chr14:35431001-38257865 | FOXA1,NKX2-1, | 5775414 | 4977134 | 7.391996 | 3 | 41 | 14 | 10 | 3 | 3 | 7 |
# 获取ecDNA 的 top20 基因
genes = paste(ecDNA_amp$OncogenesAmplified[1:nrow(ecDNA_amp)],collapse = ",") %>% str_split(pattern = ",")
genes = genes[[1]]
top20 = rev(head(tail(sort(table(genes)),n=21),n=20))
top20_gene = names(top20)
# top20 gene 对应的 amp 类型
amp_top20 = data.frame()
for (i in top20_gene) {
amp_gene = amp[grep(pattern = i,ignore.case = F,x = amp$OncogenesAmplified),]
amp_gene$gene = i
amp_top20 = rbind(amp_top20,amp_gene)
}
# 柱状图可视化
amp_top20$gene = factor(amp_top20$gene,levels = top20_gene)
amp_top20$class = factor(amp_top20$class,levels = c("Linear",
"Heavily rearranged",
"BFB",
"Circular"))
p = ggplot(data = amp_top20) +
geom_bar( aes(x = gene, fill = class),
#width = 0.5,
#position =position_dodge2(padding = 0.5, preserve = "single"),
stat = "count") +
# facet_grid(. ~ Patient, scales = 'free_x', space = 'free') +
theme_classic() +
theme(
panel.border = element_blank()) +
xlab(label = "top20 gene")+
ylab(label = "Frequency") +
scale_fill_manual(values = c("#377EB8", "#4DAF4A", "#FF7F00", "#984EA3"))
p
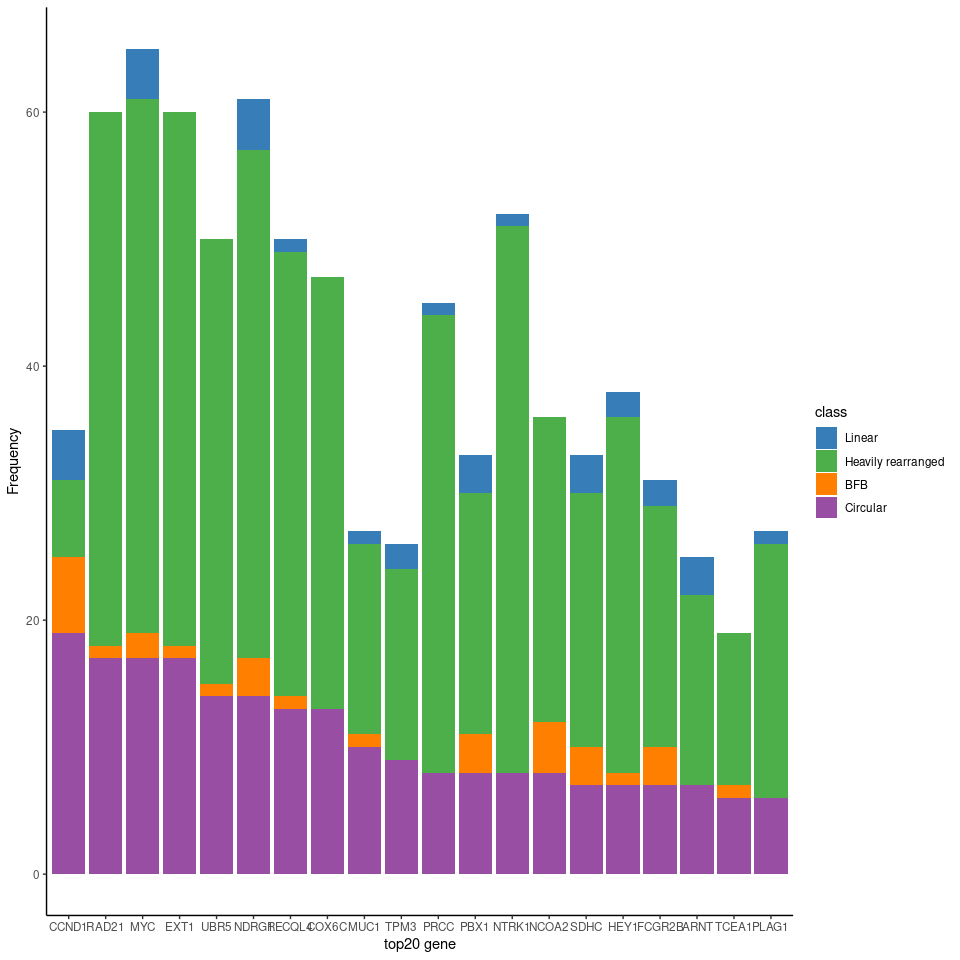
还有就是,统计出来的 top20 基因列表和文章的不一致:
top20_paper = c("CCND1","EXT1","MYC","RAD21","NDRG1",
"UBR5","COX6C","RECQL4","MUC1","TPM3",
"NCOA2","NTRK1","PBX1","PRCC","ARNT",
"FCGR2B","HEY1","SDHC","CHCHD7","MET")
amp.list = list(top20_gene=top20_gene,top20_paper=top20_paper)
library(ggvenn)
ggvenn(amp.list,
show_elements = F,
show_percentage = T,
label_sep = "\n",
fill_color = c("#E64B35B2", "#4DBBD5B2"),
auto_scale = T
)

ggvenn(amp.list,
show_elements = T,
show_percentage = F,
label_sep = "\n",
fill_color = c("#E64B35B2", "#4DBBD5B2"),
auto_scale = T
)
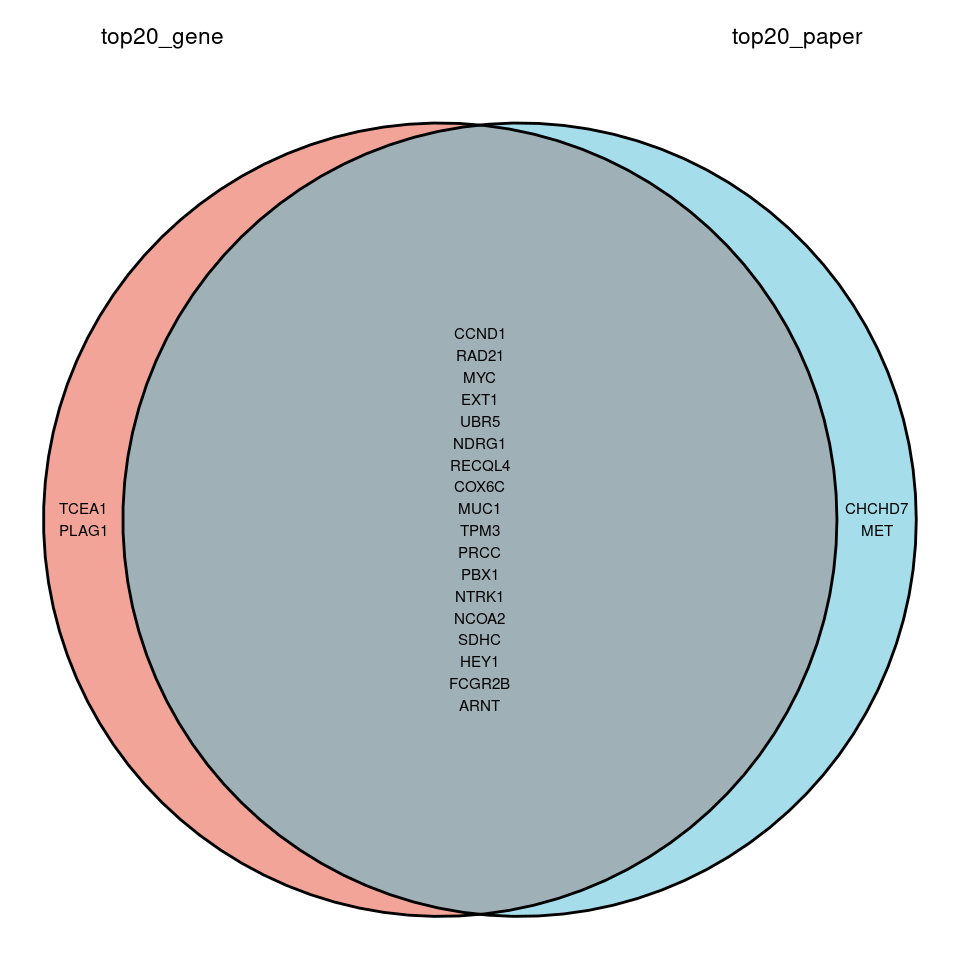
基因组重排的 circle plot
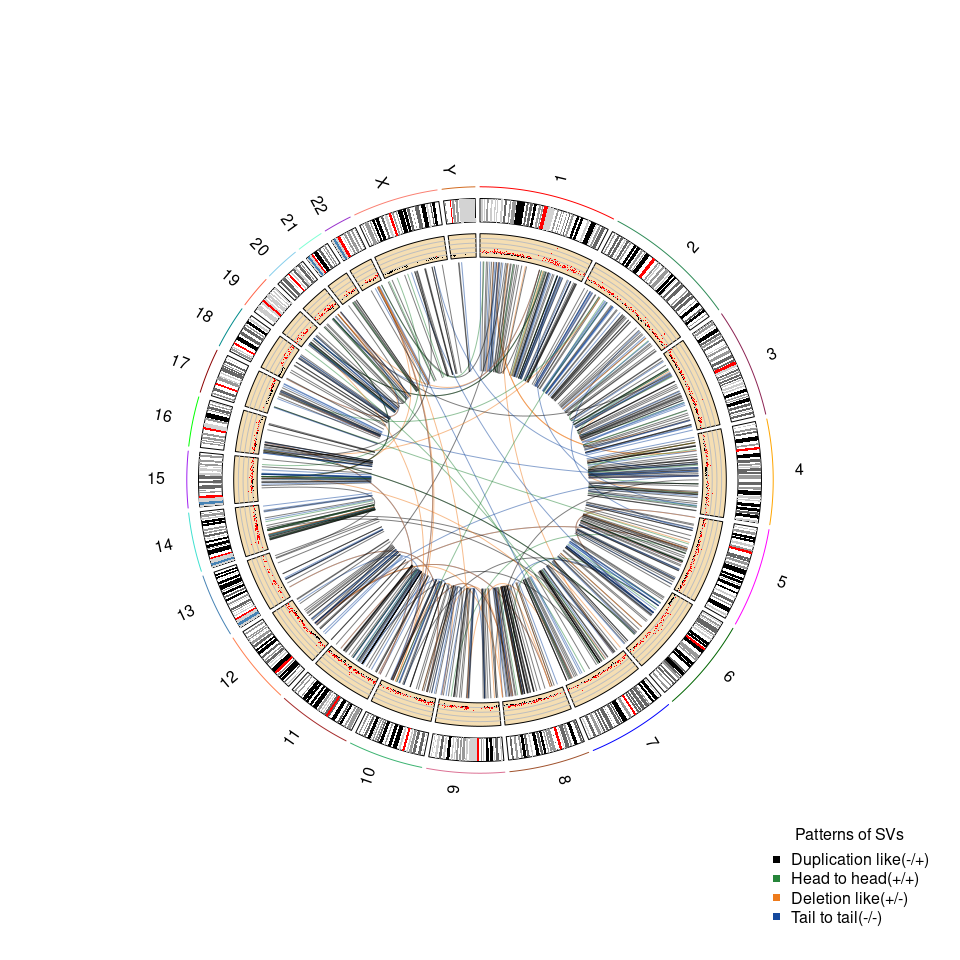
文章中的 fig.4b 是基因组重排的 circle plot,以 CLCA_0119 患者为例,circle plot 纳入了拷贝数变异信息 CN 和结构变异SV信息。

这部分信息可以从该文章报导的数据库上http://lifeome.net:8080/clca 获取到
# 读入 CN 数据
CN_data = readxl::read_xlsx("Copy_Number_Alteration_20240315.xlsx")
CN_data = as.data.frame(CN_data)
CN_data$Start = as.numeric(CN_data$Start)
CN_data$End = as.numeric(CN_data$End)
CN_data$CopyNumber = as.numeric(CN_data$CopyNumber)
# 读入 SV 数据
SV_data = readxl::read_xlsx("Structure_Variation_20240315.xlsx")
SV_data = as.data.frame(SV_data)
SV_data$PosA = as.integer(SV_data$PosA)
SV_data$PosB = as.integer(SV_data$PosB)
# 这里仔细查看发现 SV 数据的RelatedGeneB(s) 和 GeneB(s).Func 两列的信息应该颠倒过来了
head(SV_data,n=20)
CaseID | ChrA | PosA | RelatedGeneA(s) | GeneA(s).Func | A.Strand | ChrB | PosB | RelatedGeneB(s) | GeneB(s).Func | B.Strand | Chromoplexy | Chromothripsis |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
CLCA_0023 | chr1 | 8875290 | RERE | intronic | + | chr1 | 8877433 | UTR5 | RERE | - | . | . |
CLCA_0023 | chr2 | 13604672 | LOC100506474;LINC00276 | intergenic | - | chr2 | 14858959 | intergenic | FAM84A;NBAS | + | . | . |
CLCA_0023 | chr3 | 68999992 | FAM19A4;EOGT | intergenic | + | chr3 | 69000212 | intergenic | FAM19A4;EOGT | - | . | . |
CLCA_0023 | chr3 | 170362147 | LOC101928583 | ncRNA_intronic | + | chr3 | 170362183 | ncRNA_intronic | LOC101928583 | - | . | . |
CLCA_0023 | chr4 | 2495362 | RNF4 | intronic | + | chr4 | 2495727 | intronic | RNF4 | - | . | . |
CLCA_0023 | chr4 | 17266283 | LINC02493;SNORA75B | intergenic | + | chr4 | 17266334 | intergenic | LINC02493;SNORA75B | - | . | . |
CLCA_0023 | chr7 | 78027496 | MAGI2 | intronic | + | chr7 | 78028666 | intronic | MAGI2 | - | . | . |
CLCA_0023 | chr8 | 64638461 | LOC102724612;LINC01289 | intergenic | - | chr8 | 64667679 | intergenic | LOC102724612;LINC01289 | + | . | . |
CLCA_0023 | chr9 | 103941193 | PLPPR1 | intronic | + | chr9 | 103941257 | intronic | PLPPR1 | - | . | . |
CLCA_0023 | chr10 | 94663880 | EXOC6 | intronic | + | chr10 | 94663934 | intronic | EXOC6 | - | . | . |
CLCA_0023 | chr12 | 110239299 | TRPV4 | intronic | - | chr12 | 110243757 | intronic | TRPV4 | + | . | . |
CLCA_0023 | chr13 | 77667696 | MYCBP2 | intronic | + | chr13 | 77683238 | intronic | MYCBP2 | - | . | . |
CLCA_0023 | chr17 | 79257653 | SLC38A10 | intronic | + | chr17 | 79257943 | intronic | SLC38A10 | - | . | . |
CLCA_0023 | chr20 | 57261314 | STX16-NPEPL1 | ncRNA_intronic | + | chr20 | 57263239 | ncRNA_intronic | STX16-NPEPL1 | - | . | . |
CLCA_0023 | chr21 | 45289099 | AGPAT3 | intronic | + | chr21 | 45290317 | intronic | AGPAT3 | - | . | . |
CLCA_0023 | chr22 | 24834833 | ADORA2A-AS1;SPECC1L-ADORA2A | ncRNA_intronic | + | chr22 | 24911204 | exonic | UPB1 | - | . | . |
CLCA_0023 | chrX | 125812403 | DCAF12L1;PRR32 | intergenic | + | chrX | 127402085 | intergenic | ACTRT1;SMARCA1 | - | . | . |
CLCA_0023 | chr6 | 46665473 | TDRD6 | intronic | - | chr17 | 56730435 | intronic | TEX14 | - | . | . |
CLCA_0023 | chr2 | 14856430 | FAM84A;NBAS | intergenic | - | chr2 | 14917781 | intergenic | FAM84A;NBAS | - | . | . |
CLCA_0023 | chr2 | 14917766 | FAM84A;NBAS | intergenic | + | chr2 | 23985393 | intronic | ATAD2B | + | . | . |
## 获取 CLCA_0119 患者的数据
CLCA_0119_CN = CN_data[CN_data$CaseID == "CLCA_0119",2:9]
CLCA_0119_SV = SV_data[SV_data$CaseID == "CLCA_0119",2:13]
# RCircos plot
library(RCircos)
data(UCSC.HG19.Human.CytoBandIdeogram)
RCircos.Set.Core.Components(cyto.info = UCSC.HG19.Human.CytoBandIdeogram,
chr.exclude=NULL,
tracks.inside =3,
tracks.outside = 0)
RCircos.List.Plot.Parameters()
RCircos.Set.Plot.Area()
RCircos.Chromosome.Ideogram.Plot()
# 添加拷贝数变异信息,散点图
RCircos.Scatter.Plot(scatter.data = CLCA_0119_CN,
data.col=4,
track.num=1,
side="in",
by.fold=2);
# 添加结构变异曲线
## 添加End,这里只是为了方便可视化,所以 End 是在 start 上加1,没有实际意义的
CLCA_0119_SV$EndA = CLCA_0119_SV$PosA+1
CLCA_0119_SV$EndB = CLCA_0119_SV$PosB+1
## 添加 Patterns 进行分类,原数据没有,但是文章的RCircos plot 有
CLCA_0119_SV$Patterns =
ifelse(CLCA_0119_SV$A.Strand == "+" & CLCA_0119_SV$B.Strand == "+",
yes = "Head to head(+/+)",
ifelse(CLCA_0119_SV$A.Strand == "-" & CLCA_0119_SV$B.Strand == "-",
yes = "Tail to tail(-/-)",
ifelse(CLCA_0119_SV$A.Strand == "+" & CLCA_0119_SV$B.Strand == "-",
yes = "Deletion like(+/-)",
no = "Duplication like(-/+)")))
## 添加 PlotColor 设置颜色
CLCA_0119_SV$PlotColor =
ifelse(CLCA_0119_SV$A.Strand == "+" & CLCA_0119_SV$B.Strand == "+",
yes = "black",
ifelse(CLCA_0119_SV$A.Strand == "-" & CLCA_0119_SV$B.Strand == "-",
yes = "#26853A",
ifelse(CLCA_0119_SV$A.Strand == "+" & CLCA_0119_SV$B.Strand == "-",
yes = "#EE7B1C",
no = "#15499D")))
## 重新进行列排序
CLCA_0119_SV_link = CLCA_0119_SV[,c("ChrA","PosA","EndA","ChrB","PosB","EndB","PlotColor",
"RelatedGeneA(s)", "GeneA(s).Func","A.Strand",
"RelatedGeneB(s)", "GeneB(s).Func","B.Strand",
"Chromoplexy","Chromothripsis","Patterns" )]
CLCA_0119_SV_link$ChrA = factor(CLCA_0119_SV_link$ChrA,levels = c(paste0("chr",c(1:22,"X","Y"))))
CLCA_0119_SV_link$ChrB = factor(CLCA_0119_SV_link$ChrB,levels = c(paste0("chr",c(1:22,"X","Y"))))
RCircos.Link.Plot(
link.data = CLCA_0119_SV_link,
track.num = 2,
# by.chromosome = T,
#start.pos = 0.8,
genomic.columns = 3,
is.sorted = T
)
legend("bottomright",
#inset=.05,
title="Patterns of SVs",
legend = c(unique(CLCA_0119_SV_link$Patterns)),
#lty=1,
pch=15, bty = "n",
col=c("black", "#26853A","#EE7B1C","#15499D"))
sessionInfo()
## R version 4.3.2 (2023-10-31)
## Platform: x86_64-pc-linux-gnu (64-bit)
## Running under: Ubuntu 20.04.6 LTS
##
## Matrix products: default
## BLAS/LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.8.so; LAPACK version 3.9.0
##
## locale:
## [1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
## [3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
## [5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
## [7] LC_PAPER=en_US.UTF-8 LC_NAME=C
## [9] LC_ADDRESS=C LC_TELEPHONE=C
## [11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
##
## time zone: Asia/Shanghai
## tzcode source: system (glibc)
##
## attached base packages:
## [1] parallel stats4 grid stats graphics grDevices utils
## [8] datasets methods base
##
## other attached packages:
## [1] RCircos_1.2.2 ggvenn_0.1.10
## [3] dplyr_1.1.4 ggstatsplot_0.12.1
## [5] purrr_1.0.2 sigminer_2.3.0
## [7] doParallel_1.0.17 iterators_1.0.14
## [9] foreach_1.5.2 BSgenome.Hsapiens.UCSC.hg19_1.4.3
## [11] BSgenome_1.70.2 rtracklayer_1.62.0
## [13] BiocIO_1.12.0 Biostrings_2.70.3
## [15] XVector_0.42.0 GenomicRanges_1.54.1
## [17] GenomeInfoDb_1.38.8 IRanges_2.36.0
## [19] S4Vectors_0.40.2 barplot3d_1.0.1
## [21] NMF_0.26 synchronicity_1.3.10
## [23] bigmemory_4.6.1 Biobase_2.62.0
## [25] BiocGenerics_0.48.1 cluster_2.1.6
## [27] rngtools_1.5.2 registry_0.5-1
## [29] ggVennDiagram_1.4.9 VennDiagram_1.7.3
## [31] futile.logger_1.4.3 ggsci_3.0.0
## [33] ggrepel_0.9.4 pheatmap_1.0.12
## [35] data.table_1.15.4 tidyr_1.3.0
## [37] ggpubr_0.6.0 ggplot2_3.5.0
## [39] stringr_1.5.1 maftools_2.18.0
##
## loaded via a namespace (and not attached):
## [1] splines_4.3.2 prismatic_1.1.1
## [3] bitops_1.0-7 ggplotify_0.1.2
## [5] tibble_3.2.1 R.oo_1.25.0
## [7] cellranger_1.1.0 datawizard_0.9.1
## [9] XML_3.99-0.16.1 lifecycle_1.0.4
## [11] rstatix_0.7.2 globals_0.16.2
## [13] lattice_0.22-5 MASS_7.3-60.0.1
## [15] insight_0.19.7 backports_1.4.1
## [17] magrittr_2.0.3 rmarkdown_2.25
## [19] yaml_2.3.8 cowplot_1.1.2
## [21] RColorBrewer_1.1-3 multcomp_1.4-25
## [23] abind_1.4-5 zlibbioc_1.48.2
## [25] R.utils_2.12.3 RCurl_1.98-1.14
## [27] yulab.utils_0.1.4 TH.data_1.1-2
## [29] sandwich_3.1-0 GenomeInfoDbData_1.2.11
## [31] correlation_0.8.4 listenv_0.9.0
## [33] parallelly_1.36.0 codetools_0.2-19
## [35] DelayedArray_0.28.0 DNAcopy_1.76.0
## [37] tidyselect_1.2.1 farver_2.1.1
## [39] matrixStats_1.2.0 GenomicAlignments_1.38.2
## [41] jsonlite_1.8.8 survival_3.5-7
## [43] emmeans_1.9.0 tools_4.3.2
## [45] rio_1.0.1 Rcpp_1.0.12
## [47] glue_1.7.0 SparseArray_1.2.4
## [49] xfun_0.42 MatrixGenerics_1.14.0
## [51] withr_3.0.0 formatR_1.14
## [53] BiocManager_1.30.22 fastmap_1.1.1
## [55] fansi_1.0.6 digest_0.6.34
## [57] R6_2.5.1 gridGraphics_0.5-1
## [59] estimability_1.4.1 colorspace_2.1-0
## [61] R.methodsS3_1.8.2 utf8_1.2.4
## [63] generics_0.1.3 S4Arrays_1.2.1
## [65] parameters_0.21.3 pkgconfig_2.0.3
## [67] gtable_0.3.4 statsExpressions_1.5.2
## [69] furrr_0.3.1 htmltools_0.5.7
## [71] carData_3.0-5 scales_1.3.0
## [73] bigmemory.sri_0.1.6 knitr_1.45
## [75] lambda.r_1.2.4 rstudioapi_0.15.0
## [77] reshape2_1.4.4 rjson_0.2.21
## [79] uuid_1.1-1 coda_0.19-4
## [81] cachem_1.0.8 zoo_1.8-12
## [83] restfulr_0.0.15 pillar_1.9.0
## [85] vctrs_0.6.5 car_3.1-2
## [87] xtable_1.8-4 paletteer_1.5.0
## [89] evaluate_0.23 zeallot_0.1.0
## [91] mvtnorm_1.2-4 cli_3.6.2
## [93] compiler_4.3.2 futile.options_1.0.1
## [95] Rsamtools_2.18.0 rlang_1.1.3
## [97] crayon_1.5.2 ggsignif_0.6.4
## [99] labeling_0.4.3 rematch2_2.1.2
## [101] plyr_1.8.9 forcats_1.0.0
## [103] fs_1.6.3 stringi_1.8.3
## [105] gridBase_0.4-7 BiocParallel_1.36.0
## [107] munsell_0.5.0 bayestestR_0.13.1
## [109] Matrix_1.6-5 patchwork_1.1.3
## [111] future_1.33.1 SummarizedExperiment_1.32.0
## [113] highr_0.10 broom_1.0.5
## [115] memoise_2.0.1 readxl_1.4.3